
ScienceAsia 33 (2007): 197-206
Dynamic Processes Permitting Stable Coexistence of
Antimicrobial Resistant and Non-Resistant Organisms in
a Gastrointestinal Tract Model
Tippawan Puttasontiphot
a
, Yongwimon Lenbury
a*
, Chontita Rattanakul
a
, Sahattaya
Rattanamongkonkul
b
, John R. Hotchkiss
c
and Philip S. Crooke
d
a
Department of Mathematics, Faculty of Science, Mahidol University,Bangkok, 10400, Thailand.
b
Department of Mathematics, Faculty of Science, Burapha University,Chonburi, 20131, Thailand.
c
Department of Critical Care Medicine, University of Pittsburgh,Pittsburgh, PA 15261, USA.
d
Biomathematics Study Group, Department of Mathematics,Vanderbilt University, Nashville, TN 37240, USA.
* Corresponding author, E-mail: [email protected]
Received 7 Jun 2006
Accepted 22 Dec 2006
ABSTRACT: In this paper, bacteria-antibiotic dynamics in a gastrointestinal tract exposed to antimicrobial
selection pressure is modeled as occurring in a continuous chemostat. Two populations of microorganisms,
one sensitive and the other resistant to an inhibitor, namely an antibiotic, compete for a single limiting
resource. It is shown that conditions exist whereby the two types of bacteria can persist. Application of the
singular perturbation analysis also yields delineating conditions on the “effective” antibiotic level that encode
information on the minimum inhibitory concentration (MIC) and the minimum antibiotic concentration
(MAC), which are extremely important parameters commonly used to quantify the activity of antibiotics
against a given bacterium.
K
EYWORDS: gastrointestinal tract model, antimicrobial resistance, continuous chemostat, singular perturbation.
doi: 10.2306/scienceasia1513-1874.2007.33.197
INTRODUCTION
As has been noted by Nolting and Derendorf
1
, “the
outcome of treating bacterial infections with antibiotics
is a function of multiple variables.” Therapeutic efficacy
depends on bacterial susceptibility to the antibiotics,
specific properties of the infected site, metabolism in
antibiotic elimination, dosing regimen, and several other
factors. Up to date, many questions about dosing
regimen remain unanswered
1
. Risk of infection may
not be reduced even though the antibiotic dose is
titrated to produce serum concentrations within the
recommended range.
Parameters traditionally used to quantify the activity
of antibiotic against bacterial infection are the minimum
inhibitory concentration (MIC), the minimum
bactericidal concentration (MBC) and the minimum
antibiotic concentration (MAC). These parameters are
determined under standardized conditions as
discussed in
Nolting and Derendorf’s report
1
. There
are several disadvantages to this approach, however.
The in vitro models do not necessarily reflect the in vivo
situation where the drug undergoes metabolism and
elimination. The inoculum size can be variable and the
infected size might pose severe restrictions on antibiotic
diffusion into the tissue
1
. Most importantly, prolonged
treatment can give rise to resistant strains. Such
concerns have prompted researchers to devise in vitro
systems or models to simulate in vivo conditions and to
investigate the killing effects of antibiotics against
bacteria as a function of time.
The mammalian digestive tract becomes colonized
with bacteria acquired, since birth, from the mother
and the immediate environment. Numerous bacterial
species populate the digestive tract and constitute the
indigeneous gastrointestinal flora of a mammal
contributing to a gastrointestinal ecosystem that is
necessary for an animal to digest and absorb nutrients
efficiently from the alimentary
2
. It is well known that
the gastrointestinal (GI) microflora that inhabit the
digestive tract of mammals have a great influence on
the host’s physiology, intestinal anatomy and resistance
to infectious diseases
3
.
Several studies
4,5
have demonstrated that
overgrowth of intestinal bacteria promotes their
translocation from the GI tract to other organs. Such
translocation of viable bacteria from the GI tract to the
mesenteric lymph nodes and other organs is
undoubtedly an initial step in the pathogenesis of many
bacterial diseases
4
. It was also reported by Steffenand
and Berg
5
that the extent of translocation of certain
indigenous bacteria is directly related to their
population levels in the GI tract.
Apparently, there is a great variety in the
198
ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
composition of gastrointestinal microflora among
animals, as well as varying degrees of sensitivity among
the microbials to the various antibiotics that might be
used to inhibit their growth. However, the emergence
of resistant microorganisms is always a potential risk
when antibiotics are administered.
Moreover, a recent report
6
indicated that there is
huge potential for genetic exchange to occur within the
dense, diverse anaerobic microbial population
inhabiting the GI tract of humans and animals.
Specifically, Scott
6
reported on recent evidence that
anaerobic bacteria native to the rumen or hindgut
harbour both novel antibiotic resistance genes and
novel conjugative transposons (CTns). These CTns,
and previously characterized CTns, can be transferred
to a wide range of commensal bacteria under laboratory
and in vivo conditions.
In their investigation of the impact of various
antibiotics on the fecal flora of newborn infants, Bennet
et al.
7
found that sequential development of the neonatal
intestinal flora can be dramatically altered upon
administration of broad-spectrum antimicrobial agents.
It takes possibly several weeks for the intestinal flora
to return to normal even when administration of the
antimicrobial agents was discontinued.
Additional evidence that demonstrates the
importance of intestinal flora in the control of GI
overgrowth was given in
the work of Burr and Sugiyama
8
, where it was shown that adult mice with normal
microbial flora but treated with large oral doses of two
broad spectrum antimicrobial agents are at least 50-
fold more susceptible to C. botulinum intestinal
colonization than untreated mice. The increased
susceptibility of antimicrobial treated mice to C.
botulinum intestinal colonization is terminated when
the antibiotics are stopped and the mice are housed
with untreated control mice.
A body of evidence
9
suggests that multiple
mechanisms are employed by the flora to exclude non-
indigeneous organisms from the intestinal tract and to
maintain the status quo. Only the most extreme stress
situations, such as antibiotic administration, have major
effects on the stability of the initial flora. Experiments
of Freter et al.
10
provided data on possible E. coli
population control mechanisms. Anaerobic continuous
flow cultures of homogenates of intestinal contents
from conventional mice mixed with E. coli in veal infusion
broth suppressed E. coli population growth, compared
with pure continuous flow cultures of E. coli. The
inhibition of E. coli multiplication was reversed,
however, by addition of glucose. These results suggested
that competition for nutrients is the overriding activity
of importance in the control of E. coli population growth
in continuous flow cultures that reflect the intestinal
conditions
10
.
Thus, in this paper, the effects of antimicrobial
selection pressure on bacterial population dynamics
and competition in the gastrointestinal tract is modeled
as occurring within a continuous chemostat. Two
populations of microorganisms in the culture compete
for a single limiting resource in the presence of a drug,
or antibiotic, as the inhibitor to which one population
is sensitive and the other resistant. Clinical data are
utilized to support the choice of terms that appear in
the model. We then investigate the conditions under
which different dynamic behavior is possible. Through
our analysis, different dosing and kill curves may be
examined. The analytical results are discussed in terms
of their clinical implications, in the hope that deeper
insights may be gained which are important for the
understanding of the stability of the microflora
composition in the GI tract, as well as the possible
factors that may disrupt such stability.
Model ConstructionModel Construction
Model ConstructionModel Construction
Model Construction
We visualize the gastrointestinal tract as a
continuous flow chemostat in which C is the
concentration of the resource, S the number of sensitive
microbial population, R that of the resistant population,
and A is the drug concentration at any moment in time.
Resource enters the habitat at concentration C
1
, while
A
1
is the asymptotic antibiotic concentration. Sensitive
and resistant populations are removed at the rates
ω
1
and
ω
2
while the resource and antibiotics are removed
at the rates
ω
3
and
ω
4
, respectively.
Experimental data have been collected from a culture
of Enterococcus faecalis ATCC 29212 and Serratia
marcescens ATCC 43861 growing in 1% Mueller-Hinton
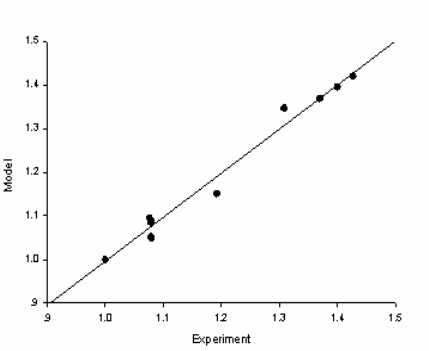
Broth II in the absence of antibiotics. The experimentally
measured values of the microbial counts are then
plotted, in Figure 1, against those obtained from the
simulation of the following logistic model for the
microbial population X(t):
Fig 1. Plot of experimental data against model simulations.

ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
199
0
()1 (0)
max
dX X
CXXX
dt X
μ
⎛⎞
=−,=
⎜⎟
⎝⎠
(1)
where
()
max
S
C
C
KC
μ
μ
=.
+
( 2 )
We observe in Figure 1 that the data points are
aligned along the straight line
y
x=
. Experimenting
with other sets of data yields the same result which
assures us that the equation (1) gives a reasonable fit
to the real data, with appropriately chosen values of the
parameters.
Next, we incorporated antibiotic effect with the
following model equation:
(3)
where A(t) = A
1
+ A
2
e
-
ω
t
( 4 )
Figure 2 shows a comparison of the simulation of
equation (3) with a set of experimental data collected
from a culture of Bacteroides Fragilis (BF) with BMS-
284756, an experimental drug, for the antibiotic, in
Mueller-Hinton Broth II. The colony counts were done
at t = 0, 2, 4, 6, 8, 24 and 48 hours by serial dilution in
normal sterile saline. The parametric values, given in
the figure caption, have been determined to obtain the
best least square fit of the kill curve. A very close fit of
this set of experimental data was obtained when the
values of the parameters were suitably chosen.
Guided by the above observations and the
discussion in the previous section, we will therefore
consider the following model system of 4 ordinary
differential equations.
Fig 2. Comparison of numerical simulation. Data generated
from equation (3), where
μ
max
=
0.02, X
max
= 2.6×10
6
,
ε
k
=
1.5945848, K
K
= 0.3911216, A
1
= 18.87638 and A
2
=
0.1,
ω
= 0.9, X
0
= 1.27×10
6
, was composed with a set of
experimental data collected from a culture of
Bacteroides Fragilis ATCC 25285 with BMS-284756 as
the antibiotic in Mueller-Hinton Broth II grown on
Tryptic-Soy agar plate.
20
2
(0)
R
R
SR
RC
dR
RR R
dt K C K S
γ
γ
ε
ψ
ω
=+−,=
++
(6)
13 0
3
() (0)
(1 )( ) ( )
SS RR
aS R
SC
RC
dC
CC C C
dt A K C K C
εψ
εψ
ω
γ
=− − − , =
++ +
(7)
14 0
1
()(0)
dA
AA A A
dt
ω
=− , =
(8)
where the first term in equation (5) is the growth
rate of the susceptible population level S, taking after
equation (2) previously mentioned. The second term
accounts for the reduction in the number of susceptible
population level as its member is converted into a
resistant strain due to the acquisition of an extra
chromosomal element, or plasmid, from members of
the resistant strain. Resistance can also be due to
chromosomal mutation that renders the strain
insensitive to the antibiotics
11
. The third term in
equation (5) is the rate at which the susceptible bacteria
is killed off by the antibiotic, taking the form suggested
by equation (3). The last term is then the rate of removal
of S by natural means. The response functions used in
equations (5) and (6) are of the Holling’s type generally
assumed in many previous population models
1,12-17
.
This means that the consumption rate increases as the
level of nutrient increases but reaches a saturation
point, beyond which rate of consumption cannot
increase even if there were more nutrient available.
Here, we do not take into account the detoxification of
the antibiotic (viz chloramphenicol by enzymatic
acetylation), assuming that this effect is negligently
small. Thus, in equation (8), the antibiotic is removed
naturally at the rate that is proportional to its amount
A at any time t.
Equation (6) describes the rate of change of the
resistant population level R growing at the rate given
by the first term which assumes a Holling type saturating
function of the substrate concentration C. To give it a
different dynamics from that of the susceptible strain,
since R is the resistant strain and thus not limited by its
environment, the growth rate of R is allowed to increase
with its number, not limited by its surrounding
conditions as in the logistic growth rate. The second
term in equation (6) accounts for the increase in R due
to the development of resistance in the susceptible
strain. It is then naturally removed at the rate
ω
2
R.
1
10
()
(1 )( )
(0)
S
aS
K
K
C
dS
SS
dt A K C
SR
AS
SS S
KSK A
γ
γ
ψ
γ
γ
ε
ε
ω
′
=−
++
−− −,=
++
(5)
0
1(0)
k
max
max K
AX
dX X
XXX
dt X K A
ε
μ
⎛⎞
=− − ,=
⎜⎟
+
⎝⎠

200
ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
The levels of S, R and C are assumed here to vary
with different time scales t
1
,
t
2
and t
3
, respectively, as
shall be explained in the next section. Equation (7)
describes the rate of change of the resource or substrate
C. The second term accounts for its consumption by
the susceptible bacteria S, while the last term accounts
for that by the resistant bacteria R. Since S is sensitive
to the presence of antibiotic A, the consumption rate
by S is reduced as A increases, and thus the factor 1+
γ
a
A
in the denominator of this term.
Since equation (8) is autonomous and depends only
on A, it can be easily solved for A(t) which is found to
eventually tend to A
1
as time progresses. We may
therefore consider the above model in the event that
A has reached the level A
1
. We are thus reduced to only
the 3 equations (5)-(7) for t ≥ t
0
, with A replaced by A
0
such that S(t
0
) = S
0
, R(t
0
) = R
0
and C(t
0
) = C
0
.
Model AnalysisModel Analysis
Model AnalysisModel Analysis
Model Analysis
In order to analyze the model equations (5)-(8) by
the singular perturbation method, we further make the
following assumptions.
As S represents the sensitive bacterial population,
it is reasonable that S should have the fastest dynamics,
while the resistant strain should be more adaptable to
the environment and less susceptible to destruction
and hence more stable, changing at a slower rate, or has
intermediate dynamics. The resource C may be kept
abundant at more or less constant level and thus
assumed to have the slowest dynamics. We therefore
utilize two small scaling parameters
ε
and
δ
, and
dimensionless variables and parameters defined as
follows.
Letting
1
0
()
102 03 0
()
St
S
tttt ttt ttxt
εεδ
=+ , = + , = + , = ,
3
2
00
()
()
() () ,
Ct
Rt
SS
yt zt=,=
1
0
1
C
S
z =,
0000
SR
K
KK
SR
SSSS
KKK
γ
γ
γ
γ
′
=, =, =,=,
and
1
1
1
S
a
A
a
ψ
γ
+
=,
1
1
2
K
K
A
KA
a
ε
+
=
1
3
1
SS
a
A
a
εψ
γ
+
,=
,
4 RR
a
εψ
= , ( 9 )
one is led to the following system.
0
()(0)
dx
fxyz x x
dt
=,,, =
(10)
0
()(0)
dy
gxyz y y
dt
ε
=,,,=
(11)
0
()(0)
dz
hxyz z z
dt
εδ
=,,,=
(12)
where
1
21
()
()
S
xy
axz x
fxyz ax x
Kz Kx
γ
γ
ε
γ
ω
−
,, = − − −
++
(13)
2
()
R
R
xy
yz
gxyz y
KzKx
γ
γ
ε
ψ
ω
,, = + −
++
(14)
34
13
()()
SR
axz ayz
hxyz z z
KzKz
ω
,, = − − −
++
(15)
Thus, for fixed x, y and z, the rate of change
dz
dt
is
O(
εδ
),
dy
dt
is O(
ε
), while
(1)
dx
dt
O=
. This means that x
has the fastest dynamics, y the intermediate, and z the
slowest dynamics. We can then carry out a singular
perturbation analysis on the model system (10)-(12)
with (13)-(15) that involves purely geometric
arguments. Such method of analysis has been developed
by Muratori and Rinaldi
12
, extended to 3-dimensional
systems by Muratori
13
. Examples of its use may be
found in the works of Muratori and Rinaldi
14
, Lenbury
and Likasiri
15
and Rattanakul et al.
16
. Through the
identification of the shapes and locations in the phase
space of the equilibrium manifolds where f(x,y,z) = 0,
g(x,y,z) = 0 and h(x,y,z) = 0, conditions can be determined
under which as time passes, the three state variables
x(t), y(t) and z(t) tend to the corresponding steady state
values x
S
, y
S
and z
S
, respectively, at which point f(x
S
,
y
S
,z
S
) = g(x
S
,y
S
,z
S
)= h(x
S
,y
S
,z
S
) = 0. The direction of the
trajectories in the phase space is determined by
considering the signs of
xy,
and in different regions
in the (x,y,z) - space delineated by the equilibrium
manifolds. We are particularly interested in the case
where the wash-out steady state (0,0,z
1
) is stable and
the bacteria populations are eradicated.
Since experimental data invariably show oscillatory
behavior in the time series, as can be seen in Figure 3,
we are also interested in the possibility that our model
allows such behavior. The conditions under which this
occurs are important for control purposes and yield
insightful interpretations.
Fig 3.
Colony counts of Serratia marcescens ATCC 43861 in
varying concentration of 0.1% MHB with ceftazidime
(0.125 ng/ml) on Trypticase-Soy agar plate. The colony
counts were done at t = 0, 1, 3, 5, 7 hours by serial
dilution in normal sterile saline. Experimental data are
shown as dots, while the solid curve corresponds to the
model simulation.
z
ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
201
The shapes and relative positions of the manifolds
{f = 0}, {g = 0}, and {h = 0} determine the directions,
speeds, and shapes of the resulting solution trajectories.
Therefore, we need to analyze each of the equilibrium
manifolds in detail. The delineating conditions for
occurrence of different dynamic behavior as well as
the existence of limit cycles are arrived at from the
close inspection of those manifolds.
The manifold The manifold
The manifold The manifold
The manifold
{{
{{
{
ff
ff
f
= 0} = 0}
= 0} = 0}
= 0}
This manifold consists of two submanifolds. One is
the trivial manifold x = 0, while the other is the nontrivial
manifold given by the equation
y =
112 12
()()
()
S
S
aK aK z aKK
Kz
γγ γ
γ
γω ω
ε
⎡⎤
⎢⎥
⎣⎦
−+ −+
+
+
2
112121
[( )( )] ( )
()
S
S
aK az aKxazx
Kz
γ
γ
γω ω
ε
⎡⎤
⎢⎥
⎣⎦
−−+ −+ −
+
(16)
The surface intersects the z-axis at the point where
12
2
112
()
()
S
aK
zz
aa
ω
γω
+
=≡
−+
(17)
In order that z
2
> 0, we need to require
112
()aa
γω
>+
(18)
Moreover, the curve where the surface intersects
the (y, z)-plane is asymptotic to the line
112
1
()aK aK
yy
γγ
γ
γω
ε
−+
=≡
(19)
for which y
1
> 0 provided (18) holds. Also the curve
where the surface intersects the (x, z)-plane is
asymptotic to the line
112
1
1
()aa
xx
a
γ
ω
−+
=≡
(20)
for which x
1
> 0 provided (18) holds.
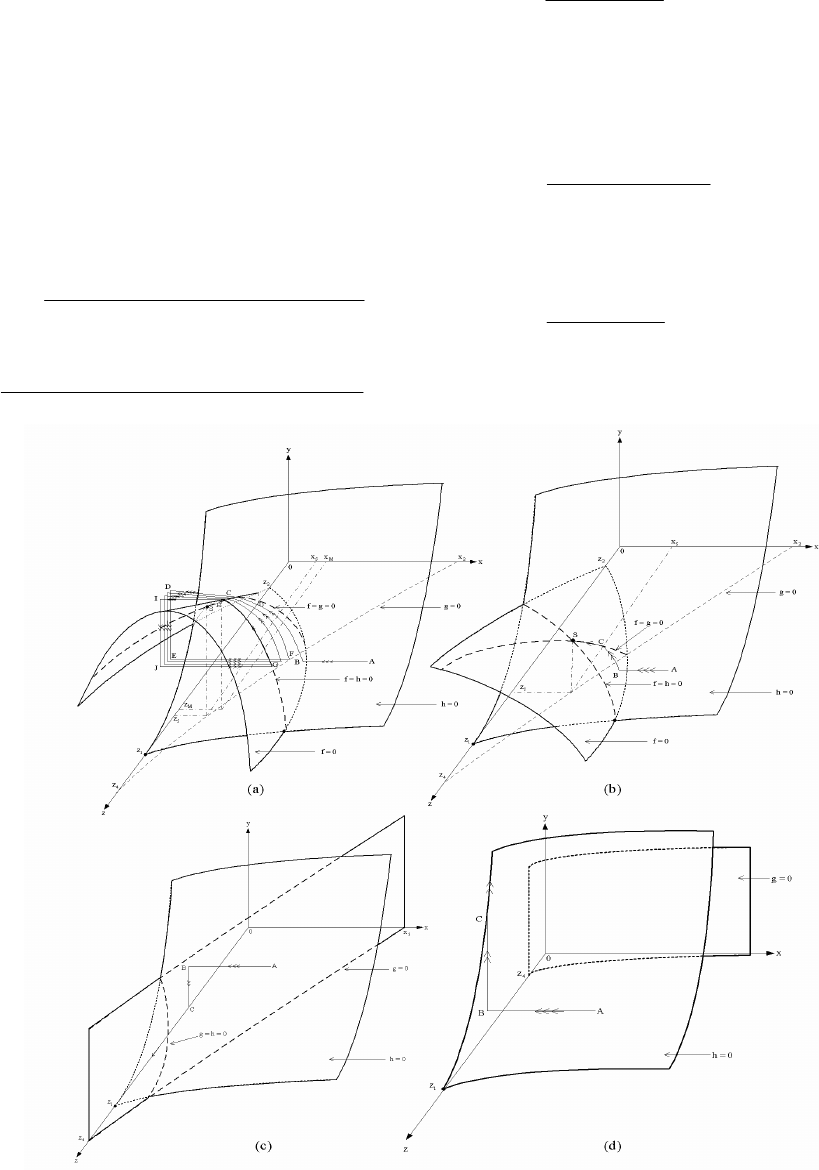
Fig 4.
Trajectories of the model system (10)-(12) with (13)-(15). The trajectory tends to a limit cycle in a), a positive equilibrium
point in b), a wash-out equilibrium point in c), and becomes unbounded in d).

202
ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
We now consider
y
x
∂
∂
, the slope of y given by (16)
as a function of x, for a fixed z, as follows.
112121
()( )( )2
()
S
S
aK az aKaxz
y
xKz
γ
γ
γω ω
ε
⎡⎤
−−+ −+ −
∂
⎣⎦
=
∂+
(21)
We see that
y
x
∂
∂
= 0 along the curve where
11212
1
()( )( )
()
2
S
max
aK az aK
xz x
az
γ
γω ω
⎡⎤
−−+ −+
⎣⎦
==
(22)
In order that x
max
> 0, we need
12
3
112
()
()( )
S
aK
zz
aK a
γ
ω
γω
+
>≡
−−+
(23)
We observe that z
3
> 0 if
112
()aK a
γ
γω
−>+
(24)
in which case the manifold {f = 0} is shaped and
located in the phase space shown in Figure 4a.
The manifold The manifold
The manifold The manifold
The manifold
{{
{{
{
gg
gg
g
= 0} = 0}
= 0} = 0}
= 0}
This manifold consists of two submanifolds. One is
the trivial manifold y = 0, while the other is the nontrivial
manifold given by the equation
22
22
()
()()
RR
RR
Kz KK
x
zK
γγ
γγ
ψω ω
ωψ ε ωε
−−
=
−− + −
(25)
This nontrivial manifold is independent of the
intermediate variable y, and thus it is parallel to the y -
axis. It intersects the z- axis at the point where
2
4
2
R
R
K
zz
ω
ψω
=≡
−
(26)
which will be positive if
2R
ψ
ω
> (27)
Moreover, it intersects the x-axis at the point where
2
2
2
K
xx
γ
γ
ω
εω
=≡
−
(28)
which will be positive if
2
γ
ωε
<
(29)
The manifold The manifold
The manifold The manifold
The manifold
{{
{{
{
hh
hh
h
= 0} = 0}
= 0} = 0}
= 0}
This manifold is given by the equation
133
4
()()() ()
()
SR R
S
zzK zK z axzK z
y
azK z
ω
−++− +
=
+
(30)
which intersects the
z
-axis (x = y = 0) at the point
where
(31)
We see that we need to require that
41
zz> (32)
and
21
xx> (33)
for the surfaces to be located as shown in Figures
4a-d.
We now let (x
S
,y
S
,z
S
) be the intersection point of {f
= 0}, {g = 0} and{h = 0}, with x
S
> 0, y
S
> 0, z
S
> 0, and let
(x
M
,y
M
,z
M
) be the maximum point of the curve f = h = 0.
Four cases of different dynamic behavior can be
identified as follows.
Case 1Case 1
Case 1Case 1
Case 1
..
..
. This case is identified by the inequalities
(18), (24), (27), (29), (32), (33), and 0 < x
S
< x
M
.
Under these conditions, the shapes and relative
positions of the equilibrium manifolds {f = 0}, {g = 0} and
{h = 0} are as depicted in Figure 4a, where three arrows
indicate fast transitions, two arrows indicate
intermediate ones, while single arrows indicate slow
ones.
Thus, a system initially at a generic point, say point
A of Figure 4a, will make a quick transition to the
surface given by equation (16) of the fast manifold {f =
0} (point B in Figure 4a), since the signs of f assure us
that this part of the manifold {f = 0} is stable. As the
surface where {f = 0} is approached, y has slowly become
active. A transition at an intermediate speed is made
along {f = 0} in the direction of increasing y, since g > 0
here.
Once the point C is reached, the stability of the
manifold is lost, a catastrophic transition brings the
system to point D on the (x,z)-plane. Consequently, the
system will slowly develop along this plane in the
direction of decreasing y, since g is now negative.
At a point E on the (y,z)-plane, the stability will again
be lost and a quick transition will bring the system back
to the point F which almost closes a cycle. However,
since this part of the transition is in the region
where
0z >
, z slowly increases giving rise to a high
frequency oscillation which densely covers a surface,
until the point H on the curve {f = h = 0} is reached. A
quick jump brings the trajectory to point I which is on
the (y,z)-plane, and an intermediate transition will bring
the system to the point J, followed by a quick transition
which will bring the system back to the point G which
lies on the stable part of the curve{f = h = 0}, thereby
forming a closed cycle GHIJG. Thus, the existence of a
limit cycle in the system for
ε
and
δ
sufficiently small
is assured.
Case 2Case 2
Case 2Case 2
Case 2. This case is identified by inequalities (18),
(27), (29), (31), (32) and (33) while (24) is violated (see
Figure 4b).
Under these conditions, the system initially at a
generic point A will make a quick transition to the fast
manifold {f = 0} (point B), then an intermediate transition
is made along this surface in the direction of increasing
1
0zz=>
ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
203
y until the curve {f = g = 0} is reached. The trajectory will
follow this curve until the stable equilibrium point
(x
S
,y
S
,z
S
) where {f = g = h = 0} is reached at which point
the transition ceases. This is, therefore, the case where
all three populations persist at constant stable levels.
Case 3Case 3
Case 3Case 3
Case 3
..
..
. This case is identified by inequalities (27),
(29), (32), and (33) while (18), and thus (24), are
violated. In Figure 4c, we see that the nontrivial fast
manifold does not appear in the first octant since (18)
does not hold. The intermediate manifold and the slow
manifold are as shown in Figure 4c. A system initially
at a point A, where the level of substrate C
0
is not too
high, will make a quick transition to the trivial manifold
x = 0, namely the (y,z)-plane (point B). Then, since g <
0 here, a transition at an intermediate speed in the
direction of decreasing y is made to the point C on the
z-axis. Then the slow variable z has become active, and
a transition at a slow speed is made along the z-axis to
end at the stable equilibrium point(0, 0, z
1
). This is thus
the case in which both susceptible and resistant bacterial
populations are washed out.
Case 4Case 4
Case 4Case 4
Case 4
..
..
. This case is identified by inequalities (27),
(29), and (33), while (18), (24), and (32) are violated.
Also, this is the case where z
1
>> z
4
(high feed substrate
or efficient substrate consumption by the resistant
population). The system initially at a point A (in Figure
4d) will make a quick transition in the direction of
decreasing x until a point B on the manifold {h = 0} is
reached (as shown in Figure 4d). At this point, the
intermediate variable y has become active and a
transition at intermediate speed is then made along {h
= 0} in the direction of increasing y, since g > 0 here in
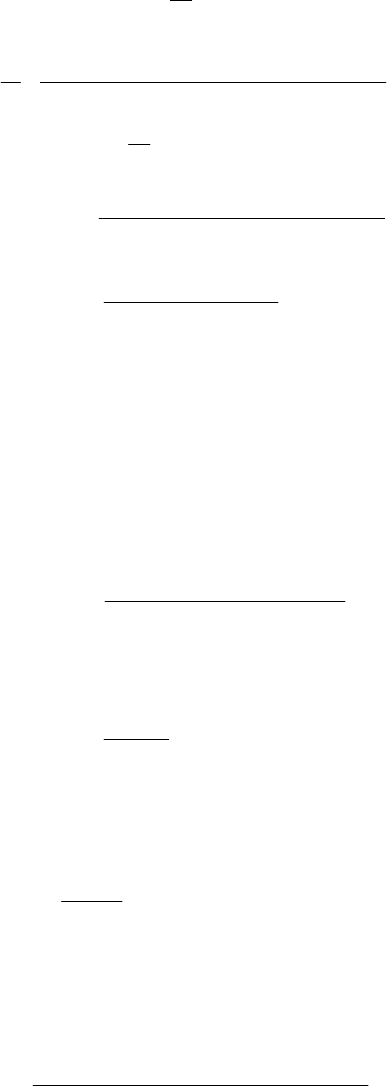
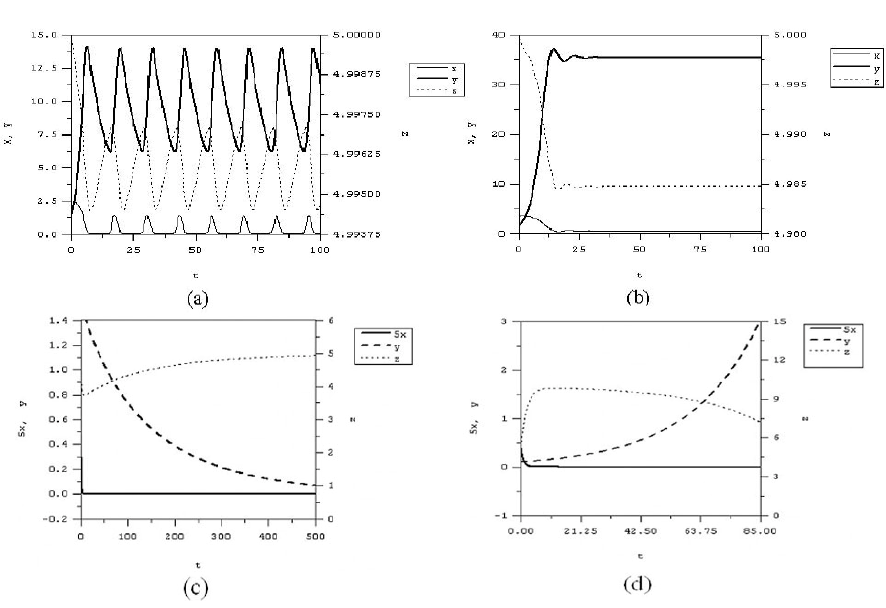
Fig 5.
Time courses of x, y and
z
corresponding to the 4 cases of Figure 4.
(a)
ε
= 1.0,
δ
= 1.0, z
1
= 5.0,
ε
γ
= 0.9,
ψ
R
= 1.2,
γ
= 3.0,
ω
1
= 0.7,
ω
2
= 0.7,
ω
3
= 0.7, K
S
= 0.7, K
γ
= 1.5, K
R
= 5.0, a
1
= 2.45283,
a
2
= 0.023077, a
3
= 0.00196, a
4
= 0.0006, x
0
= 0.5, y
0
= 1.5, and z
0
= 5.0. The solution in Case 1 exhibits sustained
oscillation.
(b)
ε
= 1.0,
δ
= 1.0, z
1
= 5.0,
ε
γ
= 0.9,
ψ
R
= 1.2,
γ
= 4.0,
ω
1
= 0.7,
ω
2
= 0.7,
ω
3
= 0.7, K
S
= 0.7, K
γ
= 4.3, K
R
= 5.0, a
1
= 2.407407,
a
2
= 0.023077, a
3
= 0.00196, a
4
= 0.0006, x
0
= 0.5, y
0
= 1.5, and z
0
= 5.0. The solution in Case 2 is seen here to tend towards
a stable non-washout steady state values.
(c)
ε
= 1.0,
δ
= 1.0, z
1
= 5.0,
ε
γ
= 0.1,
ψ
R
= 0.09291,
γ
= 2.0,
ω
1
= 0.5,
ω
2
= 0.09,
ω
3
= 0.5, K
S
= 2.0, K
γ
= 2.0, K
R
= 0.5, a
1
= 0.1, a
2
= 0.8, a
3
= 0.5, a
4
= 0.5, x
0
= 0.5, y
0
= 1.5, and z
0
= 5.0. The solution in Case 3 is seen here to tend towards a stable
wash-out steady state values.
(d)
ε
= 1.0,
δ
= 1.0, z
1
= 10.0,
ε
γ
= 0.06,
ψ
R
= 0.1,
γ
= 2.0,
ω
1
= 0.5,
ω
2
= 0.06,
ω
3
= 0.5, K
S
= 2.0, K
γ
= 2.0, K
R
= 0.00001,
a
1
= 0.1, a
2
= 0.8, a
3
= 0.5, a
4
= 0.5, x
0
= 0.1, y
0
= 0.1, and z
0
= 5.0. The solution in Case 4 is seen here to exhibit a steady
increase in the resistant population y.

204
ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
front of the manifold {g = 0}, until the point C on the
curve {h = 0} on the plane {x = 0} is reached. The
transition will continue along this curve so that the
resistant population can become too high leading to a
bacteria overgrowth in the GI tract. The results of the
above analysis may be stated as in the following theorem.
TheorTheor
TheorTheor
Theor
emem
emem
em For the model system (10)-(12), with (13)-
(15), and a suitable starting point(x
0
,y
0
,z
0
) , suppose
inequalities (27), (29) and (33) hold.
1. If inequalities (18), (24) and (32) are satisfied and
0 < x
S
< x
M
, then there exists a periodic solution to the
model system which is locally asymptotically stable.
The solution trajectory starting close enough to the
unstable non-washout steady state (x
S
,y
S
,z
S
) in this case
will tend, as time progresses, towards a limit cycle
which surrounds that steady state.
2. If the inequality (24) is violated, while (18) and
(32) still hold, then the non-washout steady state will
be stable. In this case, we have coexistence of the two
microbial species at steady state levels.
3. If, moreover, (18) is violated (in which case (24)
also is), while (32) still holds, then the washout steady
state (x,y,z) = (0, 0, z
1
) is locally stable. In this case, there
is an
ε
0
> 0 such that if C
0
<
ε
0
, then bacteria are eventually
eradicated.
4. On the other hand, when the environment is too
conductive to bacteria growth, there exists a such
that if C
0
>
ˆ
C
and (18), (24), and (32) are violated, with
z
1
>> z
4
, the mechanism to control the resistant strain
through nutrient competition is no longer viable. The
antibiotic dose of A
0
which settles down eventually to
the level A
1
in this case is high enough to violate (24).
The susceptible population is therefore quickly
eliminated, allowing the overgrowth of the resistant
population.
Numerical Simulation and InterpretationNumerical Simulation and Interpretation
Numerical Simulation and InterpretationNumerical Simulation and Interpretation
Numerical Simulation and Interpretation
Figure 5 shows the time series of the numerical
simulations of model equations (10)-(12), with (13)-
(15). The parameters are chosen to satisfy (18), (24),
(27), (29), (32) and (33) in Figure 5a yielding a limit
cycle solution as theoretically predicted. The solution
trajectory in Figure 5b tends to the non-washout
equilibrium point, while in Figure 5c it tends to the
washout steady state (0, 0, z
1
) if it starts with a low C
0
.
However, if C
0
is too high bacteria overgrowth may be
expected as shown in Figure 5d. We observe here that
although our analysis was done on the assumption of
ε
and
δ
being small in order to carry out the singular
perturbation arguments, our conclusions are in fact
still valid even without
ε
or
δ
being very small. Both are
set equal to 1 in the simulations shown in Figure 5.
In order to understand the clinical implications of
the conditions given in the different cases of the above
theorem, we consider the delineating conditions (18)
and (24). We define
2
a
as the value of a
2
that satisfies
(18) when inequality is replaced by equality, namely,
1
2
a
a
γω
≡− (34)
and
2
a
as the value of a
2
that satisfies (24) when
inequality is replaced by equality, namely,
1
2
aK
a
γ
γω
≡−−
(35)
The above analytical conclusions may be
understood to say that if the antibiotic dose can be kept
at the sustaining level A
1
, low enough so that the
“inhibition factor” a
2
remains below the level
2
a
, but
kept high enough so that a
2
exceeds the critical level
2
a
,
then we shall be in Case 2 of the above theorem. That
is, if
2
22
a
a
a
<<
(36)
then the two bacterial populations can persist at
controllable stable steady state levels (Case 2). However,
if the antibiotic level asymptotically settles toward the
level A
1
that is high enough so that the inhibition factor
a
2
exceeds both
2
a
and
2
a
, given in (34) and (35), then
the inequalities (18) and (24) are both violated. We may
then expect an eventual eradication of both the sensitive
and the resistant populations (Case 3), provided that
the initial condition (substrate level C
0
) is not too highly
conductive to bacteria growth.
Figure 6 shows different “kill curves” from our
model simulations for different “effective” antibiotic
levels A
1
. The oscillatory behavior exhibited by our
model in these kill curves closely resembles that
Fig 6.
Kill curves for different “effective” antibiotic levels.
ε
=
1.0,
δ
= 1.0, z
1
= 5.0,
ε
γ
= 0.9,
ψ
R
= 0.5,
γ
′′
′′
′
= 2.5,
γ
a
= 1.0,
ω
1
= 0.5,
ω
2
= 0.7,
ω
3
= 0.5, K
S
= 2.0, K
γ
= 2.0, K
R
= 0.5,
K
K
= 1.0,
ε
S
= 0.000814,
ε
R
= 0.0006,
ε
K
= 0.046154,
ψ
S
= 4.814814, x
0
= 0.5, y
0
= 1.5, z
0
= 5.0 and S
0
= 1.0, A
1
= antibiotic level.
1
1
ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
205
observed in clinical data, an example of which is shown
in Figure 3 where a simulation of our model is shown
fitted with the experimental data, taking into account
the measurement errors. Solving (34) for A
1
gives us an
indication of the MAC (minimum antibiotic
concentration), while solving (35) for A
1
provides a
lower bound for the MIC (minimum inhibitory
concentration). According to Sharma et al
18
the amount
of time during which the antibacterial concentration
remains above the MIC or MAC is an important
determinant of deciding the dose regimens of an
antibacterial agent.
However, the “effective” antibiotic level A
1
for each
patient cannot be measured, but may by obtained by
interpolating the pharmaco-kinetic data collected from
controlled clinical trials. Considering equation (8), if
the antibiotic rate of change of A
is plotted against the
antibiotic level A at each moment in time, then the slope
of the fitted line that best fits the data shall give the value
of
ω
4
, and the y-intersect of the line gives the value of
A
1
ω
4
, thus yielding the effective antibiotic level A
1
characteristic to each patient. In this way, it is possible
to design the dosing regimen to obtain the desired
outcome in the control range given by (36).
Finally, in terms of maintenance of the community
stability in the GI tract, we can offer the following
interpretation. The transition from a stable situation of
Case 2 seen in Figure 5b to the unstable situation of
Case 1 seen in Figure 5a occurs with the realization of
condition (24). Considering this condition, we see that
it may be satisfied with low enough antibiotic level A
1
,
or sufficiently low removal rate of susceptible
population
ω
1
, or high enough carrying capacity
γ
of
the environment for the susceptible population, or
faster conversion of susceptible to resistant strain (low
K
γ
), or high enough initial nutrient level C
0
.
Also, transition from the stable situation of Case 3
seen in Figure 6c to that seen in Figure 5d again is
facilitated by the abundance of nutrients (large z
1
) or
efficient consumption of nutrients by the resistant
population (high
ψ
R
or low K
R
) so that z
1
>> z
4
even with
a high level of antimicrobial agents A
1
. Our model thus
bears support to the experimental observation of Freter
et al.
10
, already mentioned in the Introduction, that
suggested that competition for nutrients is of an
overriding importance in the activity that maintains the
stability of the microflora community. From such
analysis, we can suggest a possible control strategy that
combines appropriate drug protocol and strict diet
regimen.
CONCLUSION
We have discussed a model of bacteria responses
and resistance to antibiotics, which accounts for the
mechanisms involved in the bacteria-antibiotic
interactions in vivo. For a specific patient, it is possible
to determine the values of the parameters by
interpolating the data collected from controlled clinical
trials, as has been described above. The model can then
be used to simulate different dynamic behavior due to
different dosing regimes. Since routine application of
antibiotics inevitably leads to the emergence of drug
resistance, it is vital that strategies are divised to reduce
the speed with which this occurs.
Clinically, it is difficult to assess pharmaco-dynamic
effects of antibiotics due to the complexity in repeatedly
determining the bacterial load at the site of infection
and antibiotic concentrations during the dosing
interval
18
. Using dynamic models of bacteria-antibiotic
interactions can overcome these difficulties. Through
the model development and analysis, we gain an
understanding of the pharmaco-dynamic factors that
are involved in the process, while our model tries to
simulate human infection mechanisms and can suggest
how best the microbiological activity and antibacterial
pharmacokinetic data in vivo can be used to select an
antimicrobial agent and its dosage regimen that
minimizes GI tract overgrowth by resistant species.
The model might also be used to provide information
regarding the kinetics of elimination of resistant strains
once antimicrobial selection pressure has been relieved.
Such information could prove useful in designing,
monitoring, and control of empiric therapy stragies.
ACKNOWLEDGEMENTS
Appreciations are extended towards the Thailand
Research Fund and National Center for Genetic
Engineering and Biotechnology, National Science and
Technology Development Agency (contract no. 3-
2548) for financial support.
REFERENCES
1. Nolting and Derendorf H (1995) Pharmacokinetic/
Pharmacodynamic Modeling of Antibiotics. CRC Press.
2. Johnson SA, Nicolson SW and Jackson S (2004) The effect
of different oral antibiotics on the gastrointestinal microflora
of a wild rodent (Aethomys namaquensis). Comparative
Biochemistry and Physiology,
A138A138
A138A138
A138, pp 475-83.
3. Tannock GW (1999) Medical importance of the normal
microflora. Kluwer Academic Publ., Dordrecht, the
Netherlands,.
4. Van der Waaij D, Berghuis-deVries JM and Lekkerkerk-van
der Wees JEC (1972) Colonization resistance of the digestive
tract and the spread of bacteria to the lymphatic organs in
mice. J. Hyg.
70,70,
70,70,
70, pp 335-42.
5. Steffenand EK and Berg RD (1983) Relationship between
cecal population levels of indigenous bacteria and
translocation to the mesenteric lymph nodes. Infection and
Immunity
39(3)39(3)
39(3)39(3)
39(3), pp 1252-9.
6. Scott KP (2002) The role of conjugative transposons in
206
ScienceAsia ScienceAsia
ScienceAsia ScienceAsia
ScienceAsia
33 (2007)33 (2007)
33 (2007)33 (2007)
33 (2007)
spreading antibiotic resistance between bacteria that inhabit
the gastrointestinal tract. Cell. Mol. Life Sci.
5959
5959
59
,,
,,
, pp 2071-82.
7. Bennet R, Erickssen M, Nord CE and Zetterstrom R (1984)
Impact of various antibiotics on the fecal flora of newborn
infants. Microecol. Ther.
1414
1414
14, pp 251.
8. Burr DH and Sugiyama H (1982) Susceptibility to enteric
botulinum colonization of antibiotic-treated adult mice.
Infect. Immun.
3636
3636
36
, ,
, ,
, pp103-6.
9. Van der Waaij D, Heidt PJ, Rusch VC and Gebbers JO (1990)
Microbial ecology of the human digestive tract. In: Old Herborn
University seminar monograph, pp 1-83. Institute for
Microecology, Germany.
10. Freter R, Abrams GD and Aranki A (1973) Patterns of
interaction in gnotobiotic mice among bacteria of a synthetic
“normal” intestinal flora. In: Germfree Research: Biological effect
of gnotobiotic environments (Ed.: Heneghan, J.B.). pp 429-
434. Academic Press, New York, London.
11. Peacock SJ, Mandal S and ICJW (2002) Bowler, Preventing
Staphylococcus aureus infection in the renal unit. Q J Med
9595
9595
95, pp 405-10.
12. Muratori S and Rinaldi S (2002) A separation condition for
the existence of limit cycles in slow-fast systems. International
report, Elettronica
3,3,
3,3,
3, pp 90.
13. Muratori S (1991) An application of the saparation principle
for detecting slow – fact limit cycle in three – dimensional
system. Appl. Math. Comput.
4343
4343
43, pp 1-18.
14. Muratori S and Rinaldi S (1992) Low – and high – frequency
oscillations in three – dimensional food chain systems. SIAM
Journal on Applied Mathematics.
52,52,
52,52,
52, pp 1688-706.
15. Lenbury Y and Likasiri C (1994) Low-and high-frequency
oscillations in a food chain where one of the competing
species feeds on the other. Math. Comp. Modelling.
20,20,
20,20,
20, pp
71-89.
16. Rattanakul C, Lenbury Y, Krishnamara N and Wollkind DJ
(2003)Modeling of bone formation and resorption mediated
by parathyroid hormone: response to estrogen/PTH therapy.
BioSystems
70,70,
70,70,
70, pp 55-72.
17. Leah EK (1988) Mathematical models in biology. Random
House, New York.
18. Sharma KK, Sangraula H and Mediratta PK (2002) Some
New Concepts in Antibacterial Drug Therapy. Indian Journal
of Pharmacology
3434
3434
34, pp 390-6.
