
Parasitology
cambridge.org/par
Review
Cite this article: Faria JRC (2021). A nuclear
enterprise: zooming in on nuclear organization
and gene expression control in the African
trypanosome. Parasitology 148,1237–1253.
https://doi.org/10.1017/S0031182020002437
Received: 25 October 2020
Revised: 22 December 2020
Accepted: 24 December 2020
First published online: 7 January 2021
Key words:
Antigenic variation; genome architecture;
nuclear bodies; RNA processing; gene
expression; transcription factories;
Trypanosoma brucei; Trypanosomatids; VSG
Author for correspondence:
Joana R. C. Faria,
E-mail: jrcorreia[email protected]
© The Author(s), 2021. Published by
Cambridge University Press. This is an Open
Access article, distributed under the terms of
the Creative Commons Attribution licence
(http://creativecommons.org/licenses/by/4.0/),
which permits unrestricted re-use,
distribution, and reproduction in any medium,
provided the origina l work is properly cited.
A nuclear enterprise: zooming in on nuclear
organizatio n and gene expression control in the
African trypanosome
Joana R. C. Faria
The Wellcome Trust Centre for Anti-Infectives Research, School of Life Sciences, University of Dundee, Dow Street,
Dundee DD1 5EH, UK
Abstract
African trypanosomes are early divergent protozoan parasites responsible for high mortality
and morbidity as well as a great economic burden among the world’s poorest populations.
Trypanosomes undergo antigenic variation in their mammalian hosts, a highly sophisticated
immune evasion mechanism. Their nuclear organization and mechanisms for gene expression
control present several conventional features but also a number of striking differences to the
mammalian counterparts. Some of these unorthodox characteristics, such as lack of controlled
transcription initiation or enhancer sequences, render their monogenic antigen transcription,
which is critical for successful antigenic variation, even more enigmatic. Recent technological
developments have advanced our understanding of nuclear organization and gene expression
control in trypanosomes, opening novel research avenues. This review is focused on
Trypanosoma brucei nuclear organization and how it impacts gene expression, with an
emphasis on antigen expression. It highlights several dedicated sub-nuclear bodies that com-
partmentalize specific functions, whilst outlining similarities and differences to more complex
eukaryotes. Notably, understanding the mechanisms underpinning antigen as well as general
gene expression control is of great importance, as it might help designing effective control
strategies against these organisms.
Introduction
Trypanosomes are members of the Euglenozoa, a group of organisms within the Incertae sedis
Eukarya (ex-Excavata) supergroup (Adl et al., 2019). These organisms are likely to have
branched very early during evolution, which may explain the vast number of unorthodox fea-
tures that define their biology (Navarro et al., 2007; Adl et al., 2019). Euglenozoa include
Euglenids, Phytomonads and Trypanosomatids, free-living phagotrophs, plant and animal
parasites, respectively (Adl et al., 2019). Trypanosomatids include several parasitic protozoa
that cause a huge health and economic burden amongst the world’s poorest populations;
these include Leishmania sp, Trypanosoma cruzi, Trypanosoma brucei, Trypanosoma congo-
lense and Trypanosoma vivax. Notably, climate change, increased mobility and mass migration
pose great challenges to our ability to control diseases caused by these organisms, rendering
the need for new drugs to fight new parasite strains and resistance emergence imperative.
Therefore, a detailed molecular understanding of fundamental aspects of their cell biology,
gene expression, metabolism and interaction with the hosts is critical to design effective con-
trol strategies.
Trypanosoma brucei is the causative agent of sleeping sickness and nagana in humans and
cattle, respectively, and has been used for decades as a model organism for this group mostly
given its genetic tractability and available tools for reverse and forward genetics (Djikeng et al.,
2001; Alsford et al., 2011; Dean et al., 2017; Rico et al., 2018).
Trypanosoma brucei is transmitted through the bite of a tsetse fly and rapidly differentiates
into ‘slender’ bloodstream forms (BSFs) in the mammalian host. The slender forms are capable
of sensing the population density, which triggers differentiation into stumpy form s. The latter
are pre-adapted to life in the tsetse, where they will eventually differentiate into the procyclic
forms. In the mammalian host, besides the BSFs, these parasites can occupy multiple tissues
(brain, adipose tissue, skin, etc.), some recently identified as important reservoirs (Capewell
et al., 2016; Trindade et al., 2016).
Mammalian-infective T. brucei undergoes antigenic variation to successfully evade the host
adaptive immune responses (Fig. 1A), similarly to other pathogens such as malaria and giar-
diasis causing parasites (Duraisingh and Horn, 2016). For that purpose, it relies on a vast gen-
etic repertoire of genes that encode for their variant surface glycoprotein (>2500 VSG genes
and pseudogenes), approximately one-third of its genome (Berriman et al., 2005; Muller
et al., 2018). There are two key features for successful antigenic variation: (1) the ability to
express a single antigen from myriad possibilities (monogenic expression); (2) the ability to
switch from one antigen isoform to another (Duraisingh and Horn, 2016). However, despite
the vast genetic repertoire, a VSG gene can only be expressed from a limited subset of sub-
telomeric transcription units known as expression-sites (ESs) (Navarro and Cross,
1996;
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
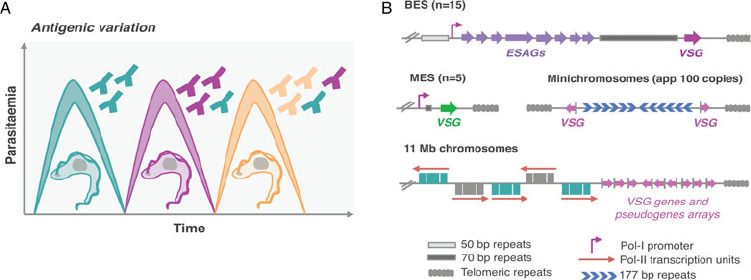
Hertz-Fowler et al., 2008 ; Fig. 1B). VSG-ESs are polycistronic
transcription units (PTUs) that share the same DNA elements,
and yet, one is active whereas the remaining are silent – a classic
epigenetic paradigm (Duraisingh and Horn, 2016).
The molecular understanding of the mechanisms underpin-
ning antigenic variation is critical as it sustains persistent infec-
tions and has greatly challenged vaccine development against
these organisms. This review will be focused on nuclear compart-
mentalization and how it affects or might affect both antigen and
global gene expression in the African trypanosome. Overall,
T. brucei nuclear architecture and mechanisms for gene expres-
sion control follow some of the classic conventions but also pre-
sent phenomenal dissimilarities when compared to so-called
model eukaryotes.
Genome organization
Eukaryotic genomes are condensed by several orders of magni-
tude; such compaction is critical to fit into the nucleus of a cell.
This is achieved by coiling the DNA around histones forming
chromatin fibres, which are subsequently arranged into more
complex high-order structures such as loops, domains and com-
partments (Gibcus and Dekker, 2013; Finn and Misteli, 2019).
Several of these architectural features are conserved across the
evolutionary tree, suggesting an elementary role of spatial organ-
ization in genome function and gene expression control (Foster
and Bridger, 2005). Indeed, DNA spatial organization and compart-
mentalization has been found to play a key role in the regulation of
gene expression and recombination in multiple organisms. In mam-
mals on a larger scale, two major sub-nuclear compartments can be
defined, one is transcription-permissive (compartment A) and the
other transcription-repressive (compartment B), roughly corre-
sponding to euchromatin and heterochromatin, respectively
(Gibcus and Dekker, 2013; Finn and Misteli, 2019). Further, within
chromatin domains known as topologically associating domains
(TADs), chromatin loops modulate interactions between promoters
and distal regulatory elements, ultimately impacting gene expression
(Rao et al., 2014; Schoenfelder and Fraser, 2019). TADs are usually
defined by boundary elements containing architectural chromatin
proteins; these include cohesin, CCCTC-binding factor (CTCF)
and histone variants (Millau and Gaudreau, 2011; Merkenschlager
and Odom, 2013).
The core genome of the African trypanosome
Trypanosoma brucei has a diploid genome, the haploid nuclear
genome (32 Mbp) is divided into three classes of linear chromo-
somes: 11 pairs of megabase chromosomes (at least 1 Mbp), one
to five intermediate-sized chromosomes and more than 100 mini-
chromosomes (50–150 kbp) (Wickstead et al., 2004; Berriman
et al., 2005). The megabase chromosomes contain all
RNA-Polymerase-II (Pol-II) transcribed genes and VSG-ESs
(Berriman et al., 2005; Muller et al., 2018). The minichromo-
somes are highly repetitive and many also contain VSG genes
(Wickstead et al., 2004; Fig. 1B). Additionally, T. brucei has an
unknown number of highly repetitive circular extra-chromosomal
DNAs of unknown function (Alsford et al., 2003).
In trypanosomes, electron-dense chromatin regions can be
found close to the nuclear periphery and their arrangement is devel-
opmentally regulated (Belli, 2000;Eliaset al.,
2001;Navarroet al.,
2007). Indeed, in T. brucei, chromosome conformational capture
(Hi-C) revealed that the transcribed chromosome core regions
and the sub-telomeric regions coding for the large reservoir of silent
VSG genes appear to fold into structurally distinct compartments
(Muller et al., 2018), similar to active A and silent B compartments
described in mammalian cells (Schoenfelder and Fraser, 2019).
Further, the relative interaction frequency was substantially higher
across sub-telomeric regions compared to core regions, indicating
that sub-telomeres are more compact than the core region.
Additionally, centromeres and junctions between the core and sub-
telomeres were found to be the most prominent boundaries of DNA
compartments (Muller et al., 2018).
Regarding architectural chromatin proteins, while CTCF
appears to be absent in non-metazoans (Heger et al., 2012), the
major subunit of cohesin is present in T. brucei and its depletion
is lethal (Landeira et al., 2009). Moreover, histone variants (H3V
and H4V) also function as architectural proteins in this organism
(Muller et al., 2018). Indeed, studies in T. brucei H3V and H4V
knockout cell lines revealed changes in global genome architec-
ture and local chromatin configuration, which triggered switches
in VSG expression (Muller et al., 2018).
The telomeres and sub-telomeres
Genome sequences of T. brucei and Plasmodium revealed that
Pol-II transcribed genes are located in the central core and
Fig. 1. Antigenic variation in T. brucei bloodstream forms. (A) Antigenic variation. Waves of parasitaemia are a hallmark of infections by African trypanosomes in
mammals. This is due to waves of parasites expressing different VSG coats (different colours). VSGs are highly immunogenic, typically triggering an effective and
lasting immune response (immunosuppression can occur later during infection). This illustration is a simplified depiction of the in vivo dynamics, indeed, at any
time point the populations can be much more complex than represented: these may include large numbers of different clonal VSG variants. (B) Genomic organ-
ization of VSG genes. Bloodstream VSG expression-sites (BESs) contain expression-site-associated genes (ESAGs), which are located between the promoter and the
70 bp repeats. The VSG genes are near telomeric repeats. Large extensions of 50 bp repeats are located upstream of all BESs. Metacyclic VSG expression-sites (MESs)
lack ESAGs and are expressed in metacyclic trypomastigotes in the salivary glands of the tsetse fly. Pol-II transcribed genes are organized in long polycistronic
transcription units in the 11 megabase (Mb) size chromosomes. The arrows indicate the direction of Pol-II transcription. VSG genes or pseudogenes are organized
in sub-telomeric regions of megabase chromosomes or at the telomeres of minichromosomes.
1238 Joana R. C. Faria
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
antigen genes are located in sub-telomeric regions (Berriman
et al., 2005; Otto et al., 2018; Fig. 1B).
The telomere is a special functional complex at the end of lin-
ear chromosomes, consisting of tandem repeat DNA sequences
and associated proteins, which can form a specialized heterochro-
matic structure that suppresses the expression of genes located at
the sub-telomere, known as telomere position effect or telomeric
silencing (Ottaviani et al., 2008). Telomeres are essential for gen-
ome integrity and chromosome stability in eukaryotes and their
synthesis is mainly achieved by the cellular reverse transcriptase
telomerase, an RNA-dependent DNA polymerase that adds telo-
meric DNA to telomeres (Cong et al., 2002). Telomerase activity
was found to be absent in most normal human somatic cells, which
is intimately related with the ageing process, but present in over 90%
of cancerous cells (Cong et al., 2002). Notably, telomere-binding
proteins play critical r oles on the maintenance of telomer e length,
telomere heterochro ma tin formation, regulation of the telomeric
transcript levels, among others (Ottaviani et al., 2008). The mam-
malian telomere comple x has been well chara cterized and contains
six core proteins that include TRF1, TRF2, TIN2, RAP1, TPP1 and
POT1 (de Lange, 2005). Additionally, an integral component of
telomeric heterochromatin is the telomeric repeat-containing RNA
(TERRA), a large non-coding RNA whose transcription occurs at
most or all chromosome ends. Further, R-Loops have been identi-
fied at the telomeres, these are three-str anded nucleic acid structures
that contain a DNA:RNA hybrid. R-Loops can play an important
role in a number of cellular functions but they can also be an
instability factor (T an and Lan, 2020).
In the insect-stage, trypanosome telomer es tend to be close to the
nuclear periphery, but this is much less pronounced in the
mammalian-s t age (DuBois et al., 2012). In T. brucei, besides the tel-
omerase components (Dr eesen et al., 2005;Sandhuet al., 2013),
which are critical for telomere maintenance, sev er al other telomere
pr oteins have been identified. Among these, TbTRF, a functional
homologue of mammalian TRF2, a TbTRF-intera cting factor,
TIF2, RAP1 and TelAP1 (Yang et al., 2009;Jehiet al., 2014a,
2014b;Reiset al., 2018). Except for TelAP1, all the other factors
are essential for cell viability; TbTRF and TbTIF2 are critical for telo-
mere integrity and their depletion leads to an increase in double-
strand breaks and increased VSG switching (Jehi et al., 2014a,
2014b). TbRAP1 intera cts with TbTRF and its depletion leads to
derepression of silent VSG
-ESs in the mammalian- infectiv e stage,
but also in insect-stage cells, where VSG expr ession is developmen-
tally shut down (Yang et al., 2009). Further, TbRAP1-mediated
silencing has a str onger impact on telomere proximal genes (Yang
et al., 2009). Moreover, by associating with telomere chromatin,
TbRAP1 also suppresses the expr ession of the TERRA transcripts
and telomeric R-Loops, consistent with a role on telomere integrity
(Nanavaty et al., 2017). Recent studies on T. brucei ribonuclease H
enzymes, endonuclease enzymes that ca taly se the cleavage of RNA
in an RNA/DNA substrate, also showed that R-loops at the telomere
and the sub-telomer e affect VSG switching frequencies (Briggs et al.,
2018, 2019).
Interestingly, the nuclear phosphatidylinositol 5-phosphatase
(PIP5Pase), part of the inositol phosphate pathway, has been
recently shown to interact with TbRAP1 in a ∼0.9-MDa complex
(Cestari et al., 2019). The inositol phosphate pathway regulates
several cellular processes in eukaryotes including chromatin
remodelling and gene expression, and had been shown to have
a role on telomere silencing and VSG monogenic expression in
T. brucei (Cestari and Stuart, 2015).
In summary, in T. brucei (similarly to Plasmodium), Pol-II
transcribed genes are located in the central core whereas the anti-
gen genes are located in sub-telomeric regions (Berriman et al.,
2005; Otto et al., 2018). This chromos ome partitioning may be
important to fine-tune recombination in regions that encode for
antigens and to ensure that all but one antigen is repressed.
Similarly to Plasmodium, there is a large amount of evidence
that supports a role for telomeric chromatin in VSG gene silen-
cing (Duraisingh and Horn, 2016). Moreover, the sub-telomeric
location of VSG-ESs is thought to favour recombination, since
these sites are rather unstable (Glover et al., 2013).
Recombination-based and transcriptional mechanisms can lead
to VSG switching, but undoubtedly recombination makes the lar-
gest contribution in T. brucei compared to Plasmodium. Indeed,
telomere integrity and stability impacts VSG swit ching frequen-
cies and has been also proven critical to maintain VSG monogenic
expression (reviewed by Saha et al.,
2020). Notably, one of the
many remaining outstanding questions is how the active
VSG-ES escapes telomeric silencing.
Remarkably, the active VSG-ES and the silent VSG-ESs reside
within distinct nuclear compartments; the importance of nuclear
compartmentalization on global gene expression control and VSG
expression, in particular, will be addressed in the next chapter.
Nuclear compartmentalization
The nucleus is a double lipid bilayer enclosed organelle, which
separates genomic DNA from the rest of the cell. Its architecture
shields the genome from the sources of damage whilst providing
opportunities for gene expression regulation (reviewed by Lin and
Hoelz, 2019). There is ample evidence in multiple eukaryotes that
the transcriptional activity of genes is influenced by nuclear
organization, which changes during differentiation and develop-
ment. Indeed, the regulated expression of genes during develop-
ment is influenced by the availability of regulatory proteins and
the accessibility of the DNA to the transcriptional machinery
(Finn and Misteli, 2019). In eukaryotes, heterochromatin, which
is highly compact, is mainly located at the nuclear periphery,
whereas the less compact euchromatin occupies a more interior
nuclear position.
Additionally, key nuclear functions such as transcription, rep-
lication or RNA processing are not homogeneously distributed
throughout the nuc leus and can be compartmentalized. Such
compartmentalization within the nucleopla sm enables functional
specialization, separation of conflicting processes as well as
increasing the concentration of specific factors at their target
point of action (Finn and Misteli, 2019).
Two main models of nuclear organization emerged in the past. A
deterministic model proposed that specific structural elements in
the nucleus assembled into a scaffold that was then used by tran-
scriptional processes, resulting in transcriptional compartmentaliza-
tion, which was independent of active processes. Chromosome
position would therefor e be maintained by intera ctions with the
scaffold (Misteli, 2007). In striking contrast, in a self-organization
model, functional sites were formed depending on the gene activa-
tion status and without the need for predefined structures; chromo-
some position would therefore be established by chromatin itself
and interactions with functional sites. Arguably, experimental data
from many model systems strongly favour self-organization models
over deterministic models. For instance, perturbing nuclear lamins,
one of the prime structural components of the nucleus, has a
modest impact on the spatial organization of transcription and
pre-mRNA splicing sites, arguing against deterministic models
(Spann et al., 1997). Conversely, perturbation of most active nuclear
processes results in rapid chromatin architectural changes, consist-
ent with self-organization models (Misteli, 2007).
The nuclear periphery
At the nuclear periphery, there is a meshwork, designated nuclear
lamina (NL), which in mamm als is composed mainly by nuclear
Parasitology 1239
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
lamins. A growing number of nuclear proteins are known to bind
lamins and are implicated in nuclear and chromatin organization,
mechanical and genome stability, cell signalling, gene regulation,
among others (Dechat et al., 2008). Notably, many molecules
must be able to traffic between the nucleus and the cytoplasm,
rendering nucleo-cytoplasmic transport absolutely critical for
cell survival. The trafficking of macromolecules in and out of
the nucleus occurs through nuclear pore complexes (NPCs)
(reviewed by Lin and Hoelz, 2019).
Nuclear pore
NPCs are massive macromolecular assemblies! In humans, each
NPC consists of ∼1000 protein subunits, designated nucleoporins,
rendering it one of the largest protein complexes in nature
(∼110 MDa). Each NPC is located in and stabilizes an ∼800
Å-wide nuclear pore, which is generated by the fusion between
the inner and outer nuclear membranes (reviewed by Lin and
Hoelz, 2019).
NPCs are critical to maintain the nuclear integrity by prevent-
ing macromolecules from freely diffu sing in or out of the nucleus.
Macromolecules smaller than ∼40 kDa can passively diffuse
through the diffusion barrier, whereas larger macromolecules
generally do not. Facilitated transport through NPCs is rapid,
adding up to hundreds to thousands of macromolecules per
second. Notably, NPCs condu ct their cargos in their native
state, allowing macromolecules to act immediately after transport,
for instance during signal transduction (reviewed by Lin and
Hoelz, 2019).
Most NPC proteins typically form a symmetric core that pos-
sesses an 8-fold rotational symmetry (nucleoporins are incorpo-
rated in multiples of eight). This symmetric core surrounds the
central transport channel and functions as the scaffold onto
which asymmetric nucleoporins attach on the cytoplasmic and
nuclear compartments to form structures known as the cytoplas-
mic filaments and nuclear basket, respectively (reviewed by Lin
and Hoelz, 2019). One inner ring that is embedded within the
nuclear envelope, and two outer rings that reside on the inner
or outer nuclear membrane generate the symmetric core itself.
The major constituent of the outer rings in the NPC is the coat
nucleoporin complex, which serves as a structural scaffold and
docking site for other nucleoporins. The nuclear basket, com-
posed of Nup153, Nup50 and Tpr, also serves as a hub for orga-
nising nuclear architecture and modulating gene transcription,
mRNA processing and export (reviewed by Lin and Hoelz, 2019).
The majority of N PC architecture appears to be conserved
throughout the Eukaryota and was already established in the
last common eukaryotic ancestor (DeGrasse et al., 2009).
However, although the proteins and complexes are rather con-
served, their arrangements can differ substantially between cells
in the same organism or even within the same cell type at the sin-
gle cell level (Ori et al., 2013). Specifically, how the NPC connects
with the lamina and mRNA transport is likely to be highly diver -
gent between different lineages (Rout et al., 2017).
Proteomics analyses of NPC-containing fractions from T. bru-
cei provided a comprehensive inventory of its nucleoporins, which
clearly share a similar fold type, domain orga nization, compos-
ition and modularity in comparison with metazoan and yeast
(DeGrasse et al., 2009). Further, an exhaustive interactome
assigned T. brucei nucleoporins to discrete NPC substructures,
which despite retaining similar protein composition also pre-
sented remarkable architectural differences (Obado et al., 2016;
illustrated in Fig. 2). Briefly, while most elements of the inner
core are conserved, multiple peripheral structures are highly dis-
similar, possibly to accommodate divergent nuclear and cytoplas-
mic functions (Obado et al., 2016). TbNPC is highly symmetric,
with asymmetry only provided by its two nuclear basket Nups
(Obado et al., 2016). Further, orthologues of cytoplasmic Nups
or mRNA remodelling factors are absent in trypanosomes.
Notably, TbNup76, likely the cytoplasm-specific Nup82/88 ortho-
logue, localizes to both faces of the NPC (Obado et al., 2016).
Overall, trypanosomes present substantial variation in the pore
membrane proteins and the absence of critical components
involved in mRNA export in fungi and animals. Additionally,
there is evidenc e supporting a Ran-dependent system for
mRNA export in trypanosomes, which suggests distinct mechan-
isms of protein and mRNA transport (Obado et al., 2016).
TbNup110 and
TbNup92, the two components of the nuclear
basket, are predicted to have predominantly coiled-coil structure
and are likely to represent the Mlp/Tpr proteins of trypanosomes
(Holden et al., 2014). Despite performing similar roles in chromo-
some segregation, TbNup92 has a restricted taxonomic distribu-
tion and appears to have a distinct evolutionary origin than
Mlp. Further, unlike Mlp, there was no evidence for a role on
the creation of transcriptional boundaries, consistent with tryp-
anosome genome organization and gene expression control
(Holden et al., 2014). However, TbNup92-knockout cells differen-
tially expressed genes associated with RNA turnover, raising the
interesting possibility that TbNup92 might associate with a par-
ticular subset of RNA-binding proteins (Holden et al., 2014).
Notably, in T. brucei as well as related organisms, a compre-
hensive analysis on whether there are changes in the NPC com-
position or structure following differentiation into different
developmental stages is yet to be performed (Rout et al., 2017);
and if such changes occur, whether those play a role in gene
expression modulation is yet to be investigated.
Nuclear lamina
In mammals, NL is a meshwork consisting of A- and B-type
lamins and lamin-associated proteins, which lines the inner
nuclear membrane. In differentiated cells, lamin expression is crit-
ical to sustain nuclear architecture, prevent abnormal blebbing of
the nuclear envelope, and position the NPCs (Dechat et al., 2008).
NL can influence transcriptional activity and interact with a wide
range of transcription factors; it is also involved in the compaction
of peripheral chromatin (Shevelyov and Ulianov, 2019).
Eukaryotic heterochromatin, which is mainly located at the
nuclear periphery, is subdivided into densely packed constitutive
heterochromatin, including pericentromeric and telomeric
chromosomal regions, and the less condensed or so-called facul-
tative heterochromatin located in chromosomal arms (Finn and
Misteli, 2019). Chromosomal regions interacting with the NL
are designated lamina-associated domains (LADs) have been
identified in a wide-range of eukaryotes, from nematodes to
humans, and contain mostly silent or weakly expressed genes
(Shevelyov and Ulianov, 2019). This supports the idea that NL
is a repressive nuclear compartment.
Lamin genes were found in metazoa but appeared to be absent
in plants and unicellular organisms. In mammals, two major
A-type lamins (lamin A and C) and two major B-type lamins
(lamin B1 and B2) have been identified and characterized
(Dechat et al., 2008). They are composed of a long central
α-helical rod domain, flanked by globular N-terminal (head)
and C-terminal (tail) domains, which self-assemble into higher-
order structures whose basic subunit is a coiled-coil dimer
(Dechat et al., 2008). Notably, aberrant lamin protein structure
or expression can lead to irregular nuclei and abnormal gene
expression. Indeed, hundreds of mutations have been identified
in human lamins and linked to diseases, collectively known as
laminopathies that include progeria and muscular dystrophies
(Dechat et al., 2008). Interestingly, examples from yeast and
plants suggest that alternative, non-lamin, molecula r systems
can construct an NL (Dechat et al., 2008).
1240 Joana R. C. Faria
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press

In T. brucei, an analog ous to vertebrate lamins, NUP-1 is a
major compo nent of the nucleoskeleton and plays a key role on
heterochromatin organization at the nuclear periphery (DuBois
et al., 2012; illustrated in Fig. 2). NUP-1 is a critical component
of a stable network at the inner face of the trypanosome nuclear
envelope, its depletion leads to abnormally shaped nuclei and dis-
rupts NPCs and chromosomes organization (DuBois et al., 2012).
NUP-1 affinity purification led to the identification of a second
coiled-coil protein, designated NUP-2. Following NUP-2 deple-
tion, NUP-1 is mislocalized and vice versa, strongly suggesting
that NUP-1 and NUP-2 form a co-depen dent network
(Maishman et al., 2016). NUP-2 knockdown leads to severe fit-
ness cost and a dramatic impact on nuclear architecture including
severe changes to the nuclear envelope and chromosomal organ-
ization. Moreover, NUP-1 and NUP-2 are conserved across trypa-
nosomes; from a structural and functional perspective, they
behave similarly to lamins (Maishman et al ., 2016).
Notably, while the active VSG-ES resides within a transcription
factory adjacent to the nucleolus (Navarro and Gull, 2001) the
silent VSG-ESs are located at the extra-nucleolar nucleoplasm
but at more peripheral locations in BSFs (Chavez et al., 1998;
Landeira and Navarro, 2007; Fig. 2B and Fig. 3). Further, all
VSG-ESs localize to the nuclear envelope and appear to form con-
stitutive heterochromatin in insect-stage cells (Landeira and
Navarro, 2007). This is consistent with the idea that the NL is a
repressive compartment. Curiously, in Plasmodium, all silenced
var genes localize in a series of clusters at the nuclear periphery,
however, the transcription of the active var gene also occurs at a
specific site at the nuclear periphery, where the activated gene
moves away from the silenced clusters (Duraisingh et al., 2005;
Freitas-Junior et al., 2005; Lemieux et al., 2013).
In T. brucei, NUP-1 plays a role on epigenetic control of devel-
opmentally regulated loci. Indeed, following NUP-1 knockdown,
megabase chromosome telomeres reposition, multiple VSG-ESs
become active, and the frequency of VSG switching increases
(DuBois et al., 2012; Rout et al., 2017). Additionally, the active
VSG-ES promoter fails to migrate to the nuclear periphery
upon differentiation, and metacyclic VSGs are derepressed in
insect-stage cells, both likely associated with the defective forma-
tion and/or maintenance of a repressive heterochromatin com-
partment (DuBois et al., 2012). Heterochromatin-based
silencing in trypanosomes involves several proteins, such as
ISWI, RAP1 and histone deacetylase (DAC) 3 (Hughes et al.,
2007; Yang et al., 2009; Wang et al., 2010), whilst histone H1
participates in maintaining condensed chromatin in silenced
regions (Povelones et al., 2012). Strikingly, T. brucei lacks
H3K9me3, a well-characterized marker for heterochromatin,
and heterochromatin-protein 1 (HP1) (Berriman et al., 2005). It
is noteworthy that the misregulation of VSG and procyclin
genes is quite modest following NUP-1 depletion (up to
10-fold) (DuBois et al., 2012); however, it demonstrates that
NUP-1 and the trypanosome NL integrate a series of possibly
multiple mechanisms that constrain the inactive VSG-ESs and
reinforce their silent state.
Membraneless nuclear bodies
Eukaryotic cells contain membraneless organelles, designated cel-
lular bodies, which compartmentalize essential biochemical reac-
tions and cellular functions. These bodies are generated by phase
separation mediated by cooperative interactions between multiva-
lent molecules (Strom and Brangwynne, 2019; Razin and
Gavrilov, 2020). Well-characterized examples of such organelles
in the nucleus are nucleoli, which are sites of rRNA biogenesis;
Cajal bodies (CB), which are assembly sites for small nuclear ribo-
nucleoproteins (RNPs); and nuclear speckles (NSs), which are
storage compartments for RNA processing factors (Strom and
Brangwynne, 2019; Razin and Gavrilov, 2020). Besides their abil-
ity to move throughout the nucleus, another fascinating feature of
several nuclear bodies is their ability to form within the nuclear
milieu without apparent support structures, again consistently
with a self-organization model. Moreover, these organelles exhibit
properties similar to liquid droplets, being able to undergo fiss ion
and fusion. In fact, mixtures of specific RNA and certain
Fig. 2. Nuclear organization and nuclear bodies. (A) The schematics represents the mammalian nucleus, the lateral boxes highlight differences in T. brucei. (B) The
schematics represents the nuclear organization and nuclear bodies in T. brucei. Trypanosome-specific compartments are highlighted, such as the bloodstream form
(BSF)-specific expression-site body (active-VSG-Expression-Site transcription; extra-nucleolar Pol-I transcription) and the Spliced Leader (SL)-array transcription
compartments.
Parasitology 1241
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
RNA-binding proteins are able to form phase-separated bodies in
vitro (Guo et al., 2019; Hondele et al., 2019).
Nucleolus
The nucleolus is likely to be the most distinctive nuclear compart-
ment, certainly the largest and the site of ribosome biogenesis
where the 45S ribosomal repeats are clustered. Indeed,
pre-rRNA transcription and processing as well as the assembly
of the 40S and 60S complexes take place in this nuclear body
(Hernandez-Verdun et al., 2010). In animals and plants, the
nucleolus presents a tripartite substructure, which can be
observed by electron microscopy. This tripartite substructure
includes fibrillar centres (FC) surrounded by a dense fibrillar
component (DFC); both embedded in the granular component,
the biggest nucleolar subdomain composed of RNP granules.
FC stores inactive rRNA genes, whereas DFC is electron dense
given the high concentration of RNPs and is involved in early
rRNA processing (Hernandez-Verdun et al., 2010).
Eukaryotic ribosomes are composed of 18S, 5.8S, 28S and 5S
rRNA subunits and approximately 80 associated proteins. The
four rRNA molecules are the main structural and catalytic com-
ponents of the ribosome. In most eukaryotes, genes encoding
for 18S, 5.8S, 28S are organized in tandem repeats, which are tran-
scribed by RNA Polymerase I (Pol-I) into a primary transcript
further processed into the mature 18S, 5.8S, 28S rRNAs
(Hernandez-Verdun et al., 2010). Transcription occurs in the
boundary between FC and DFC. 5S rRNA genes, on the other
hand, are transcribed in the nucleoplasm by Pol-III
(Hernandez-Verdun et al., 2010).
Similarly to other eukaryotes, the nucleolus is the most dis-
tinctive membraneless sub-nuclear body in trypanosomes and
Leishmania parasites that can be easily observed by light and elec-
tron microscopy (Ogbadoyi et al., 2000; Nepomuceno-Mej ía
et al., 2010). Presently, FCs have not been identified in the nucle-
olus of these organisms, which presents a bipartite structure, simi-
larly to other protozoa, yeast, invertebrates, fish and amphibians
(Ogbadoyi et al., 2000; Nepomuceno-Mejía et al., 2010; illustrated
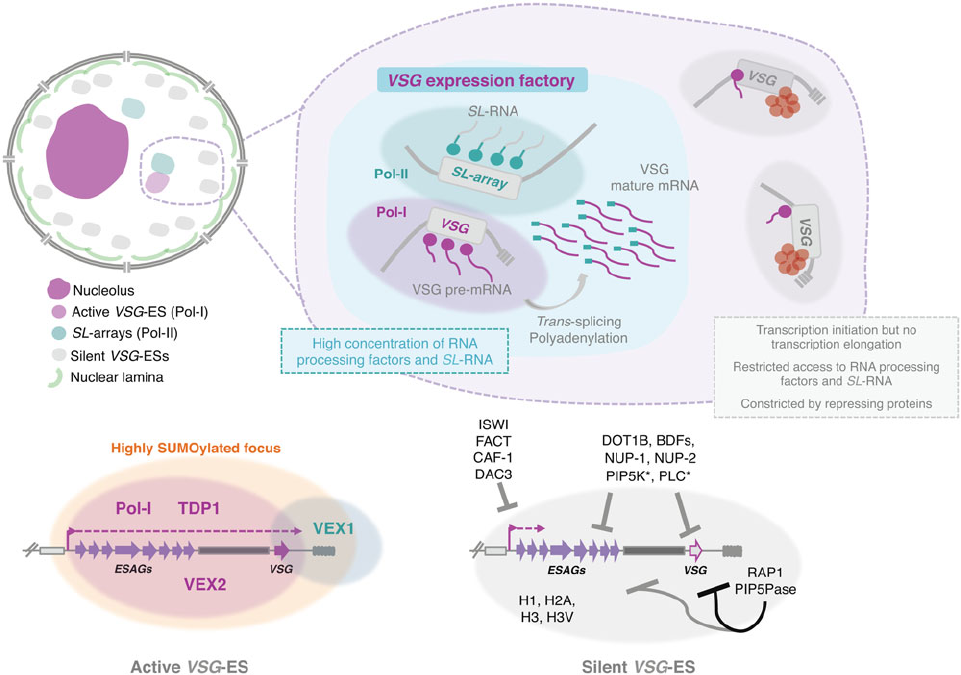
Fig. 3. Nuclear organization and VSG expression in T. brucei bloodstream forms. The single active-VSG establishes a stable inter-chromosomal interaction with one
of the SL-arrays. VEX2 orchestrates this spatial integration, which is critical to (1) sustain monog enic expression, (2) enhance RNA processing (Faria et al., 2020). The
active-VSG gene is transcribed at very high levels by Pol-I generating the most abundant protein in the cell. Proximity to the SL-array likely leads to a high local
concentration of SL-RNA therefore facilitating trans-splicing. It is possible that several factors associated with RNA processing (splicing, polyadenylation, etc .) are
concentrated in this sub-nuclear compartment as well. The SL-array appears to function as a post-transcriptional enhancer and such control might extend beyond
VSG genes (Faria et al., 2020). The active VSG-ES lies within a highly SUMOylated focus (López-Farfán et al., 2014); TDP1 is a high mobility group box protein that
facilitates Pol-I transcription and is enriched at the active-ES (Narayanan and Rudenko, 2013 ). VEX2 and VEX1 form discrete protein condensates that associate with
the active-VSG and the SL-array, respectively. The VEX complex, especially VEX2, sustains the exclusive interaction between a single VSG-ES and the SL-array; fol-
lowing its depletion, all VSG-ESs can access the SL-arrays and are derepressed (Faria et al., 2019, 2020). The silent VSG-ESs have more peripheral locations; tran-
scription by Pol-I is initiated at the same rate as at the active-locus but transcription elongation is unsuccessful; these sites also have restricted access to RNA
processing factors and substrates (Vanhamme et al., 2000; Kassem et al., 2014); several repressing factors associated with heterochromatin formation (red circles)
sustain their inactive state. For instance, ISWI (Hughes et al., 2007), FACT (Denninger and Rudenko, 2014), CAF-1 (Alsford and Horn, 2012) or DAC3 (Wang et al., 2010)
repress transcription near the promoter in silent ESs. Telomeric ES proteins, such as RAP1 (Yang et al., 2009) or PIP5Pase (Cestari et al., 2019), repress transcription
of the whole ES, the repressive gradient is stronger near the telomeres (indicated by the darker line). Other repressive proteins include DOT1B (Figueiredo et al
.,
2008), bromodomain proteins (BDFs) (Schulz et al., 2015) and PIP5K and PLC, marked with an asterisk because they are the only ones that do not localize to the
nucleus (Cestari and Stuart, 2015). Moreover, the integrity of the nuclear lamina is critical to maintain this repressive state (DuBois et al., 2012).
1242 Joana R. C. Faria
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
in Fig. 2). Unlike more complex eukaryotes, during cell division in
T. brucei, the nuclear envelope is preserved, chromatin does not
condense and the nucleolus does not disassemble. As mitosis pro-
gresses, the nucleolus stretches, is pulled via the spindle fibres to
opposite poles of the nucleus and ultimately divided into two
independent structures (Ogbadoyi et al., 2000). This process
occurs in the absence of intermediate structures such as prenu-
cleolar bodies, found in other organisms (Hernandez-Verdun
et al., 2010).
In T. brucei and similarly to other organisms, the biogenesis of
ribosome subunits starts in the nucleolus and ends in the cyto-
plasm. The 5S rRNA is imported to the nucleolus very early in
the biogenesis process and incorporated into the 90S pre-
ribosome as an RNP complex; it later undergoes spatial rearrange-
ment to facilitate subsequent maturation steps of the 60S subunit
(Prohaska and Williams, 2009; Liu et al., 2016). The pre-60S par-
ticle is translocated from the nucleus to the cytoplasm through
interactions between P34 and P37and exportin 1 and Nmd3, as
well as r-proteins uL3 and uL11 (Prohaska and Williams, 2009).
The biogenesis of the 40S subunit in T. brucei occurs very similar
to what has been described in yeast (Ferreira-Cerca et al., 2007).
Interestingly, this subunit contains a trypanosomatid-specific hel-
ical structure that has been proposed to participate in translation
initiation by interacting with the SL-sequence and its unusually
modified cap (Hashem et al., 2013).
In humans, the nucleolus has been associated with multiple
functions that extend beyond ribosome biogenesis, one being a
cellular stress sensor (Rubbi and Milner, 2003). Studies in trypa-
nosomes suggest this may be the case in trypanosomatid parasites
as well (Elias et al., 2001; Barquilla et al., 2008). Moreover, the
nucleolus appears as a largely self-organized structure. Indeed,
its integrity relies on both active Pol-I transcription and high
interactivity between ribosomal components (Raska et al.,
2006). Interestingly, ectopic expression of rRNA leads to the
formation of micronucleoli in Drosophila (Karpen et al., 1988),
again consistently with a model of self-organized nuclear com-
partmentalization. In trypanosomes, specifically, depletion of
Pol-I-specific subunits leads to abnormal nucleoli (Devaux
et al., 2007) and depletion of TOR1 kinase leads to Pol-I and
nucleolar dispersion, most likely as a consequence of Pol-I tran-
scription inhibition (Barquilla et al., 2008 ). In T. cruzi, develop-
ment from a proliferative to non-proliferative stage, which is
associated with a pronounced drop in transcriptional activity, is
also accompanied by nucleolar dispersion (Elias et al., 2001).
Further details on nucleolar structure and function in trypano-
somatid parasites have been recently reviewed by (Martínez-
Calvillo et al., 2019).
Nuclear speckles and Cajal bodies
In complex eukaryotes such as animals and plants, CBs are
involved in the post-transcriptional maturation of small nuclear
(snRNAs) and small nucleolar RNAs (snoRNAs) and the biogen-
esis of nuclear RNPs, including some nucleolar proteins,
snoRNPs and snRNPs (Sawyer et al., 2016). The number of
CBs varies across cell types and at a single-cell level within the
same cell type (in mammalian cells typically 0–10 CBs per
nucleus, ranging 0.1–2 μm in diameter). CBs are more abundant
in cells with high transcriptional activity and are highly dynamic
but structurally stable structures. They continuously exchange
components into and out of the domain in response to changes
in the cellular environmental (Sawyer et al., 2016). Interesting ly,
components of the SNAPc complex were reported to be enriched
within the CB, suggesting a strong link between snRNA gene tran-
scription and CBs. Several studies also indicate that CBs influence
the levels and processivity of factors crucial for efficient RNA
splicing; indeed CBs may influence splicing kinetics through dif-
ferent pathways (Sawyer et al., 2016).
Coilin and the nucleolar protein Nopp140 are the two key
markers of CBs. Coilin has been implicated in the link between
the nucleolus and CBs; indeed CBs are frequently detected at
the nucleolar periphery and even within nucleoli. Coilin is a key
structural component of CBs, is involved in RNP metabolism
within these nuclear bodies and it also appears to have a role
on general chromatin organization (Machyna et al., 2015). Its
N-terminal domain is responsible for the self-oligomerization
activity, truncation or mutation of phosphorylation sites in the
conserved C-terminal region leads to a dramatic alteration in
the number of CBs (Shpargel et al., 2003). On the other hand,
Nopp140 does not localize strictly to CBs and it appears to
serve generally as a chaperone for RNPs; it moves between the
nucleolus and the CBs, but also between the nucleolus and the
cytoplasm (Isaac et al., 1998). Indeed, it not only inte racts with
coilin, but also associates with several nucleolar proteins (Isaac
et al., 1998).
Trypanosoma brucei appears to lack a coilin homologue and
TbNopp140 is strictly nucleolar, strongly suggesting that CBs, in
the strict sense of the definition, are absent in these parasites
(Berriman et al., 2005; Kelly et al., 2006)(Fig. 2). Additionally,
T. brucei possesses two homologues of Nopp140, a canonical
Nopp140 and a Nopp140-like protein, both are phosphorylated
and co-immunoprecipitate with Pol-I and might play a role on
nucleoplasmic snoRNPs shuttling (Kelly et al., 2006). Given the
absence of CBs, it has been proposed that in T. brucei RNPs are
probably assembled in analogous bodies: a possible candidate
was a compartment identified as Spliced-leader-associated RNA
(SLA1)-containing subnuclear site that did not colocalize with
SL-RNA (Hury et al., 2009). SLA1 guides the pseudouridylation
at position −12 (relative to the 5
′
splice site) of the SL-RNA in
all trypanosomatid species.
NSs or splicing speckles were originally discovered as sites for
splicing factor storage and modification and were later revealed to
play a general role in RNA metabolism. Subsequently, numerous
proteins involved in epigenetic regulation, chromatin organiza-
tion, DNA repair and RNA modifications were found in NSs
(Galganski et al., 2017). Similar to other membraneless bodies
with liquid-like properties, NSs are characterized by the dynamic
exchange of components within the nucleoplasm, sharing some
proteins with other nuclear bodies (Galganski et al., 2017).
In trypanosomes, trans-splicing occurs for every single mRNA,
there are only two known cis-spliced introns in T. brucei; both
mechanisms seem to require the spliceosome. Notably, trypano-
somes encode for all snRNA and many spliceosomal proteins
described in other eukaryotes but also encode for a few specific
factors (Palfi et al., 2000; Ambrósio et al., 2009; Preusser et al.,
2009; also reviewed by Günzl, 2010; Michaeli, 2011 and
Clayton, 2019). Interestingly, splicing factors such as Prp31,
SmE, SSm2-1, PRP19 and SPF27 have a speckl e-like organization
and appear to be compartmentalized in specific nuclear areas
(Liang et al., 2006; Tkacz et al., 2007; 2010; Ambrósio et al.,
2015; illustrated in Fig. 2B).
Recent advances in more complex eukaryotes suggested that
NSs facilitate integrated regulation of gene expression
(Galganski et al., 2017). A substantial fraction of the mammalian
genome is preferentially organized around nuclear bodies such as
the nucleolus and NSs; these bodies have been proposed to act as
inter-chromosomal hubs that shape the overall packaging of DNA
in the nucleus (Quinodoz et al., 2018). Additionally, many active
genes reproducibly position near NSs, but the nature of such asso-
ciations had remained unclear until recently, when a study linked
them to stochastic gene expression amplification (Kim et al.,
2020). Whether similar associations are present and play a role
Parasitology 1243
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
in genome organization and gene expression in trypanosomes and
related organisms remains to be explored.
In summary, compartmentalization within the nucleoplasm
enables functional specialization; in fact, key nuclear functions
such as transcription or RNA processing are not homogeneously
distributed throughout the nucleus. In the next chapter, I will spe-
cifically cover the current knowledge on transcription regulation
and compartmentalization in trypanosomes.
Transcription regulation
To our knowledge, all trypanosomatids employ primarily polycis-
tronic transcription, where multiple open reading frames with no
functional association are transcribed in tandem. Evidence sug-
gests that the posi tion within the PTU is associated with messen-
ger RNA (mRNA) copy number (Kelly et al., 2012). The nascent
RNAs are processed into mature mRNAs, through a combination
of trans-splicing and polyadenylation (reviewed by Günzl, 2010;
Michaeli, 2011; Clayton, 2019). Notably, mature mRNAs bear
an unusual hypermethylated 5
′
cap structure (Bangs et al.,
1992). The genome is therefore constitutively transcribed and
mRNA abundance is primarily controlled at the post-
transcriptional level in striking contrast with more complex
eukaryotes, where a specific promoter usually regulates the tran-
scription of each gene (Koumandou et al., 2008).
Exceptions to this mechanism are the genomic loci encoding
for highly abundant surface-exposed antigens, VSGs and procy-
clins: these loci are transcribed at very high levels by Pol-I and
not Pol-II, which transcribes the majority of PTUs. Both VSGs
and procyclins expression is developmentally regulated, the for-
mer expressed in the mammalian-stage and the latter in the
insect-stage (Navarro et al., 2007; Daniels et al., 2010).
RNA polymerase I (Pol-I)
Trypanosoma brucei is the only organism known to have evolved
a multifunctional Pol-I system that is used for rRNA synthesis
and for the expression of highly abundant antigens (Günzl
et al., 2003). As previously mentioned, VSGs and procyclins are
strongly developmentally regulated and therefore Pol-I transcrip-
tion in T. brucei is intimately linked to differentiation between dif-
ferent life cycle stages as well as antigenic variation in the
mammalian host, which is critical to sustain persistent infections.
In the vast majority of eukaryotes, Pol-I is recruited to simple
promoters, which contain an upstream element located 100 bp
from the transcription start site. Such promoters are exclusively
used for rRNA gene expression, specifically the 45S rRNA precur-
sor, further processed into 18S, 5.8S and 28S rRNA. Two protein
complexes, the selectivity factor 1 (SL1) and the upstream binding
factor (UBF), are essential for Pol-I recruitment to the rRNA pro-
moter (Russell and Zomerdijk, ). The interaction between Pol-I
and SL1 is mediated by a single polypeptide named RRN3 in
humans; a UBF dimer is further required to activate rRNA tran-
scription. In yeast, RRN3 is conserved, whereas the three subunits
of the core factor (the functional equivalent of SL1) and the six
UBF subunits share no sequence similarity with the mammalian
counterparts (Russell and Zomerdijk, ). Recent CryoEM studies
suggest that, unlike the Pol-II system, promoter specificity relies
on a distinct ‘bendability’ and ‘meltability’ of the promoter
sequence that enables contacts between initiation factors, DNA
and polymerase (Engel et al., 2017). In eukaryotic cells, although
the number of rRNA genes is much lower than the number of
protein-coding genes, Pol-I transcription usually accounts for
more than 50% of the total transcriptional activity, which results
from impressively high transcription initiation rates. Notably,
mammalian Pol-I is unable to synthesize functional mRNA
(Russell and Zomerdijk, ).
Similarly to all eukaryotes, T. brucei Pol-I transcribes the 45S
rRNA precursor in the nucleolus; however, it also transcribes pro-
cyclins and VSGs mRNAs from perinucleolar and extra-nucleolar
locations, respectively (Navarro and Gull, 2001)(Figs 2B and 3).
The rRNA, VSG and procyclin gene promoters are structurally
different, suggesting that they recruit different transcription fac-
tors. Since the last two promoters are absent in related organisms
T. cruzi and Leishmania spp., one would expect to find T.
brucei-specific proteins for VSG and procyclin gene transcription.
In T. brucei, both bioinformatics and biochemical analyses
have unravelled 10 out of 12 Pol-I subunits: RPA1, RPA2,
RPC40, RPB5z, RPB6z, RPB8, RPC19, RPB10z and RPA12
(Walgraffe et al., 2005; Nguyen et al., 2006). RPB5z, RPB6z and
RPB10z are RPB5, RPB6 and RPB10 paralogs, respectively.
Further, T. brucei presents functional diversification of isoforms
that are conventionally shared RNA polymerase subunits
(Devaux et al.,
2007). Trypanosoma brucei also has a specific
component (or a divergent orthologue of yeast RPA43), RPA31,
which is critical for Pol-I transcription and cell viability
(Walgraffe et al., 2005; Nguyen et al., 2007). Although conceiv-
able, there is no evidence that RPA31, RPB5z, RPB6z and
RPB10z play a specific role on mRNA transcription by Pol-I in
T. brucei.
The class I transcription factor A (CITFA) has been identified
in T. brucei; its purification led to the identification of seven novel
subunits, termed CITFA-1 to -7, plus the dynein light chain
DYNLL1 (also known as LC8) (Brandenburg et al., 2007;
Nguyen et al., 2012). CITFA binds rRNA, VSG and procyclin pro-
moters and therefore is a general Pol-I transcription factor in T.
brucei; its depletion is unsurprising ly lethal (Brandenburg et al.,
2007). Further, TDP1, a high motility group box containing pro-
tein, which facilitates Pol-I transcription, is highly enriched at the
active VSG -ES (compared to silent) and in the nucleolus; a block-
ade in TDP1 synthesis results in a pronounced reduction of
Pol-I-derived transcripts (Narayanan and Rudenko, 2013).
TDP1 overexpression was sufficient to open the chromatin of
silent VSG-ESs and disrupt VSG monogenic expression
(Aresta-Branco et al., 2019). Moreover, ELP3B was identified as
a specific negative regulator of rRNA transcription (no impact
on VSG transcription); these observations extend the roles of
the Elp3-related proteins to Pol-I transcription units, as they are
usually associated with Pol-II transcription in humans and yeast
(Alsford and Horn, 2011).
Notably, all proteins involved in T. brucei Pol-I transcription
identified so far are conserved among all trypanosomatids, sug-
gesting that they fulfil general Pol-I functions (Walgraffe et al.,
2005; Nguyen et al., 2006; Devaux et al., 2007). However, it is
entirely possible that these common factors evolved specific func-
tions for protein-coding gene transcription in T. brucei.
Nevertheless, how T. brucei Pol-I acquired the ability to transcribe
mRNA remains mysterious.
RNA polymerase II (Pol-II)
Pol-II synthesizes pre-mRNAs and U-rich short nuclear RNAs
(snRNAs). The latter form the core of the spliceosome, involved
in processing pre-mRNAs into mature mRNAs. In both cases,
the 5
′
ends are capped, which requires adding m
7
G to the 5
′
tri-
phosphate end of the primary transcript and takes place
co-transcriptionally (Proudfoot et al., 2002). There are several
other co-transcriptional activities, which are assigned to specific
subunits or domains within these subunits (Proudfoot et al.,
2002). Trypanosoma brucei Pol-II produces both pre-mRNAs
1244 Joana R. C. Faria
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
and the spliced leader (SL)-RNA, the latter is detrimental for
trans-splicing.
Eukaryotic Pol-II enzymes usually contain 12 subunits, desig-
nated RPB1 to RPB12. Specifically, RPB1, RPB2, RPB3 and
RPB11 are considered the functional and structural core subunits.
Additionally, RPB4 to RPB10 and RPB12 usually contribute to
Pol-II ability to respond to activators and tightly bind promoter
regions (Proudfoot et al., 2002). The 12 Pol-II subunits could
be identified in T. brucei; RPB1, RPB2, RPB3 and RPB11 were
also considered the functional and structural core (Das et al.,
2006; Devaux et al., 2006). Interestingly, trypanosomes have two
isoforms of R PB5 and RPB6 (Das et al., 2006; Devaux et al.,
2006). RPB1 is the large st subunit in the T. brucei enzyme and
also the most fascinating (Evers et al., 1989). One of the most
remarkable characteristics is the non-structured carboxyl end of
the polypeptide, which deviates from the heptapeptide repeat of
YSPTSPS of varying length that is characteristic of yeast and
mammalian proteins. This repeat is generally involved in the
modulation of multiple co-transcriptional processes that include
capping, splicing, elongation, polyadenylation and nuclear export,
through coordinated kinetic alterations in the phosphorylation of
its serines and threonines (Proudfoot et al., 2002). Despite being
non-repetitive, the trypanosome carboxyterminal is phosphory-
lated and essential for transcription (Evers et al., 1989).
Trypanosoma brucei cdc2-related kinase 9 (CRK9) was found to
be responsible for RPB1 phosphorylation, however, surprisingly,
when silencing CRK9, there was no impact on Pol-II transcription
or co-transcriptional m
7
G capping. Instead it led to a block of
trans-splicing caused by hypomethylation of the SL-RNA unique
cap4 (Badjatia et al., 2013).
In many organisms, a crucial regulatory point of gene expres-
sion is transcription initiation, which requires the formation of a
pre-initiation complex that includes multiple proteins that inter-
act with Pol-II. Such transcription factors include TFIIA, TFIIB,
TFIID, TFIIE, TFIIF and TFIIH, which recruit and position
Pol-II at promoter sequences (Hahn, 2004). The only canonical
Pol-II promoter in T. brucei is the SL-RNA promoter. In this organ-
ism, the identification of general transcription factors was chal-
lenged by their extremely divergent amino acid sequences from
those of their eukaryotic counterparts. The first transcription factor
purified and characterized was a trimeric SNAPc that formed a lar-
ger complex with TATA-binding protein, the small subunit of
TFIIA (TFIIA2), and a sixth protein (TFIIA1) (Das et al., 2005;
Schimanski et al., 2005). This was followed by the identification
of TFIIB, TFIIH, TFIIE; later, a TFIIH-associated complex of
nine subunits was discovered, and despite exhibiting no motif or
sequence conservation that could reveal its identity, it structurally
resembled the head module of the much larger mediator complex
of other eukaryotes (Schimanski et al., 2006;Leeet al., 2007,
2010). More recently, a TFIIF-like or TFL complex has been iden-
tified, strongly indicating that trypanosomatids possess a full set of
RNA Pol-II general transcription factors, only very divergent from
their mammalian and yeast counterparts (Srivastava et al., 2018).
All these factors are required for SL-RNA transcription and tryp-
anosome viability, but their role, if any, on the transcription of
protein-coding genes remains unknown.
In T. brucei, ubiquitously expressed genes lack well-defined
Pol-II promoter motifs, with the exception of the spliced-leader
RNA promoter. Indeed, the so-called Pol-II disperse promoters
lack conserved sequence motifs and tight regulation; however,
they are defined by specific chromatin structures. In T. brucei
for instance, GT-rich promoters were recently proposed to drive
transcription and promote the targeted deposition of the histone
variant H2A.Z, showing that even highly dispersed, unregulated
promoters might contain specific DNA elements that are able
to induce transcription (Wedel et al., 2017).
Additionally, Pol-II transcription termination is a tightly
regulated process and critical to prevent the elongating Pol-II
complex from interfering with the transcription of downstream
genes. In kinetoplastid flagellates, the modified base
β-D-glycosyl-hydroxymethyluracil (J) replaces a small percentage
of thymine residues, mostly in telomeric regions and is synthe-
sized at the DNA level via the precursor 5-hydroxymethyluracil.
In T. brucei for instance, base J is exclusively present in the
BSF. Notably, in T. brucei and Leishmania major, base J and
H3.V are enriched at sites involved in Pol-II termination. Loss
of base J and H3.V led to transcription read-through (Reynolds
et al., 2016; Schulz et al., 2016). Recently, a novel base
J-binding protein complex involved in Pol-II transcription ter-
mination has been identified (Kieft et al., 2020).
Overall, trypanosomes appear to have limited control over
Pol-II transcription initiation, and therefore most of the gene
expression control is thought to be post-transcriptional.
RNA polymerase III (Pol-III)
Pol-III is responsible for the transcription of a number of small
non-coding RNAs that play a role in translation (tRNA and 5S
rRNAs) and other cellular processes (7SL RNA). In T. brucei,
tRNA genes can be found widely spread throughout large direc-
tional gene clusters on megabase chromosomes, 5S rRNA genes
are clustered in chromosome 8 (Berriman et al.,
2005).
Expression factories
The SL-RNA expression factory
Given that SL-RNA must be added to the 5
′
end of every single
mRNA in T. brucei, trans-splicing relies on large quantities of
SL transcripts generated by Pol-II transcription from a diploid
tandem-repeat locus. Indeed, Pol-II largest subunit is highly con-
centrated at the SL-RNA genomic loci (illustrated in Fig. 2B). In
T. cruzi and Leishmania tarentolae, a single focus is observed pos-
sibly due to pairing of both alleles (Dossin and Schenkman,
2005). In contrast, in T. brucei, two distinct foci could be detected
in G1 cells indicating that the two SL-arrays occupy distinct
chromosome territories (Uzureau et al., 2008). In T. cruzi, the
Pol-II focus disperses following treatment with transcription inhi-
bitors (Dossin and Schenkman, 2005), suggesting that the high
concentration and organization of Pol-II around the SL-arrays
depends on active-transcription and therefore is not a predefined
nuclear structure.
Moreover, SL-RNA transcripts concentrate in a nuclear area
that colocalizes with the snRNP protein SmE and SLA1 RNA,
an RNA involved in the SL-RNA modification. This strongly sug-
gests that there is a spatially defined SL-RNP factory in the
nucleoplasm (Tkacz et al., 2007). When labelling active transcrip-
tion through BrdU incorporation, a broader distribution of extra-
nucleolar transcriptional activity can be observed apart from the
SL-RNA arrays (although that accounts for Pol-III as well)
(Daniels et al., 2010; illustrated in Fig. 2B). One would expect
that capping enzymes and cap methyltransferases would concen-
trate at SL-RNP factories, which is difficult to extrapolate from the
localization data currently available and will therefore require a
more detailed analysis, possibly with higher resolution
microscopy.
The VSG expression factory
African trypanosomes and their VSGs are a fine example of
extreme biology and have led to several groundbreaking discover -
ies, such as trans-splicing, mRNA transcription by Pol-I or GPI
anchors (Navarro et al., 2007; Duraisingh and Horn, 2016).
Notably, recent studies on VSG expression in T. brucei have
Parasitology 1245
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
revealed interesting features regarding genome architecture and
nuclear compartmentalization that hint to unknown layers of
gene expression control in these organisms.
The single active VSG gene generates the most abundant pro-
tein in the cell (approximately 10% of the total proteome), which
results from a combination of high levels of transcription by Pol-I
and multiple mechanisms of post-transcriptional control
(Navarro and Gull, 2001; Günzl et al., 2003; do Nascimento
et al., 2020; Viegas et al., 2020). This renders trypanosomes and
their VSGs an amenable model system to study mechanisms
underpinning single gene choice, which are not fully understood
in any eukaryote. Indeed, monogenic expression is one of the
greatest outstanding mysterie s of eukaryotic gene expression.
For instance, it also underpins singular expression of antigen
and olfactory receptors, responsible for the specificity of the
immune response and the sense of smell in mammals, respect-
ively (Monahan and Lomvardas, 2015; Outters et al., 2015).
Interestingly, it was unclear whether genome architecture and
specifically genome position played a role in gene expression con-
trol in trypanosomes and related organisms. However, T. brucei
somehow employs a mechanism of monogenic antigen transcrip-
tion in the absence of controlled transcription initiation and
canonical enhancer sequences. Indeed, Pol-I transcription is
initiated at the same rate at all VSG-ESs, however transcription
elongation is restricted to the active-VSG-ES (Vanhamme et al.,
2000; Kassem et al., 2014). Additionally, RNA maturation
seems to be somehow restricted to the active VSG-ES suggesting
that access to RNA processing factors or substrates might be limit-
ing (Vanhamme et al., 2000; Kassem et al., 2014).
Notably, while the silent VSG-ESs were located at more per-
ipheral locations (Chaves et al., 1998; Landeira and Navarro,
2007), the active VSG-ES was included within an extra-nucleolar
structure (although in close proximity to the nucleolus), desig-
nated the expression-sit e body (ESB), a transcription factory
that contains a local reservoir of Pol-I (Navarro and Gull, 2001)
(Fig. 3). This exclusion from the nucleolus is independent of
the promoter, as swapping the VSG promoter by an rRNA pro-
moter did not lead to nucleolar incorporation (Chaves et al.,
1998), suggesting that other DNA elements/factors are required
for targeting.
The ESB emerged as the def ining structure that sustained VSG
monogenic expression, accommodating a single VSG-ES at a time.
In fact, if two VSGs were simultaneously active, a dynamic colo-
calization with the ESB was observed (Chaves et al., 1999; Budzak
et al
., 2019). However, the mechanisms for targeting the
active-VSG ES to the ESB as well as the protein composition
and the exact DNA sequences incorporated within this structure
have rem ained elusive. Although the complete molecular under-
standing is yet to be achieved, several major advances have
taken place in the recent years.
Notably, the single active-VSG displays a specific inter-
chromosomal interaction with a major mRNA splicing locus,
one of the SL-RNA arrays, and this specific nuclear arrangement
is critica l to sustain VSG monogenic expression (Faria et al.,
2020). Specifically, the single active-VSG is expressed within a
dedicated sub-nuclear compartment harbouring the Pol-I tran-
scribed antigen-coding gene and the Pol-II transcribed SL-array
and their respective associated factors to ensure (1) monogenic
antigen transcription and (2) efficient mRNA splicing (Faria
et al., 2020)(Fig. 3). The VSG exclusion proteins 1 and 2
(VEX1 and VEX2), which form discrete protein condensates in
the nucleus in BSFs specifically, associate with the SL-RNA
array and the active-VSG ES, respectively (Faria et al., 2020)
(Fig. 3). VEX1 was identified through a genetic screening
(Glover et al., 2016) and VEX2 through VEX1 affinity purification
(Faria et al., 2019). From the two proteins, VEX2, an
RNA-helicase, has the most critical role on VSG monogenic
expression: following its depletion, the ESB collapses and trypano-
somes simultaneously transcribe all VSG-ESs, subsequently
exposing multiple VSGs on their surface (Faria et al., 2019).
Further, following VEX2 knockdown, all VSG-ESs can access
the SL-RNA arrays, showing that VEX2 somehow sustains this
dedicated sub-nuclear compartment and an exclusive association
between the single active-VSG and the SL-array (Faria et al.,
2020). Additionally, besides maintaining an exclusive interaction
between the active-VSG and the SL-array, VEX2 appears to fine-
tune gene expression at the active-VSG locus (Faria et al., 2019,
2020). It is tempting to speculate that it orchestrates a specific
chromatin configuration that maximizes the interaction between
the VSG gene itself (not the promoter or the ES-associated
genes) and the
SL-array.
Phase separation and transcriptional control
More recently, liquid–liquid phase separation (LLPS) has been
proposed (opinion piece by Hnisz et al., 2017) and later demon-
strated (Guo et al., 2019) to be a major regulatory mechanism for
enhancer-mediated transcriptional control in mammalian cells.
Enhancers are short (50–1500 bp) DNA regulatory elements
that activate the transcription of specific genes to a much higher
level than would be the case in their absence; they function as a
platform for the recruitment of activators, transcription factors
and the RNA polymerase components. These DNA elements
have a distal location and are brought in proximity to the target
gene through chromatin loops. Notably, nucleation of phase-
separated multi-molecular assemblies at enhancer sequences can
explain the formation of super-enhancers (clusters of enhancers;
sometimes hundreds), their high sensitivity to transcription
inhibition, enhancer-mediated patterns of transcriptional bursts
and simultaneous activation of multiple genes by the same enhan-
cer (Hnisz et al., 2017). Notably, computational simulations have
shown that LLPS can explain experimental observations that trad-
itional models for transcriptional control cannot (Hnisz et al.,
2017).
Enhancer sequences have never been found in trypanosomes
and related parasites, and given their polycistronic transcription
and overall lack of controlled transcription initiation, such
mechanisms were thought to be unlikely to operate. But is this
really the case? Indeed, it was unclear whether and how genome
architecture and genome position played a role in gene expression
in these parasites. Could it be that trypanosomes evolved uncon-
ventional enhancers? This will be addressed in the ‘Discussion’
section.
Discussion
Despite the many open questions, previous studies following the
depletion of several chromatin-associated factors (reviewed by
Cestari and Stuart, 2018) and the recently unveiled association
between the active-VSG and the SL-array unequivocally demon-
strate that genome architecture does play a role in VSG mono-
genic transcription in T. brucei. Further, spatial proximity to
RNA-processing centres might be a conserved mechanism for
post-transcriptional enhancement of gene expression but this
had not previously been linked to inter-chromosomal
interactions.
It is possible that all VSG-ESs are able to stochastically interact
with the SL-arrays and compete for a limited pool of VEX2, which
will then stabilize an exclusive interaction between a single VSG
locus and the SL-array. This would render VEX2 a limiting factor,
which is supported by its low abundance and tight regulation
(Faria et al., 2019). Interestingly, this could be explained by an
1246 Joana R. C. Faria
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
LLPS model (Hnisz et al., 2017;Guoet al., 2019); indeed, phas e-
separating proteins were shown to be capable of generating stable
sub-nuclear structures from dynamic interactions in mammals
(Shin et al., 2018). Multiple studies have shown that high local
concentrations of specific proteins and nucleic acids (where
RNAs appear to be major players) and cooperative interactions
among these molecules are implicated in the formation of phase-
separated bodies (Shin et al., 2018; Guo et al., 2019). Recently, a
family of RNA helicases has been identified as major regulators
of the assembly of sub-nuclear compartments through LLPS
(Hondele et al., 2019); therefore, it is tempting to speculate this
might be the case of VEX2. In fact, specific post-translational
modifications can trigger nucleation of phase-separated bodies;
curiously, the active-ES resides within a hot spot of highly
SUMOylated proteins (López-Farfán et al., 2014). Notably, the
global role, if any, of LLPS and phase-separating proteins on gen-
ome organization in Trypanosomatids is yet to be investigated.
Inter-chromosomal interactions were thought to have a sto-
chastic nature, indeed the existence of stable inter-chromosomal
interactions has been a subject of debate as they were thought
to be difficult to re-establish following cell division, possibly rely-
ing on error-prone mechanisms (Finn and Misteli, 2019).
Consequently, their role on gene expression was rather dubious.
The only other known stable interaction occurs in a terminally
differentiated cell, and very interestingly, in another system sub-
ject to allelic exclusion. Indeed, olfactory neurons possess a multi-
chromosomal super-enhancer that associates with the single
active olfactory receptor gene (Monahan et al., 2019). In trypano-
somes, the association of the active-VSG with the SL-array
appears reminiscent, but classic transcriptional enhancement
was replaced by what appears to be post-transcriptional enhance-
ment instead. Despite the attractive theoretical reasons for the
presence of such an enhancer in malaria-causing parasites,
Hi-C analysis was unable to identify such an element in the P. fal-
ciparum genome (Lemieux et al., 2013).
In trypanosomes, proximity to the SL-array is likely to provide
post-transcriptional enhancement due to a high local concentra-
tion of SL-RNA. A substantial amount of SL-RNA is therefore
hijacked, so that RNA processing can keep pace with the high
rate of transcription provided by Pol-I (Fig. 3). Notably, it will
be interesting to identify other active VSG-ES-associated factors
that take part in this antigen expression factory: it is entirely con-
ceivable that a number of splicing factors and enzymes involved
in polyadenylation might be concentrated in this compartment.
This certainly adds a layer of post-transcriptional control that
had not been previously characterized. Moreover, this association
is also reminiscent of those between highly transcribed chromo-
some regions and NSs in mammals (Quinodoz et al., 2018; Kim
et al., 2020). Whether the high transcription rate is the cause or
a consequence of such association remains debatable. Similarly
in T. brucei, whether the association with the SL-RNA array pre-
cedes the activation of the VSG locus, or whether it occu rs after-
wards merely providing post-transcriptional enhancement,
remains unclear. Notably, in other organisms, co-transcriptional
RNA processing can affect transcription elongation rates
(Kornblihtt et al., 2004). How a specific VSG gene is activated
over the other possible alleles remains a mystery, and those
early events underpinning the establishment of an active tran-
scriptional state are incredibly difficult to capture. In
Plasmodium, for instance, antisense long-non-coding-RNAs
play a key role in regulating
var gene activation and mutually
exclusive expression (Amit-Avraham et al., 2015).
Certainly several mechanisms simultaneously operate to con-
strain the inactive VSG-ESs and prevent their derepression. For
instance, heterochromatin-based silencing in trypanosomes
involves, among others, ISWI, RAP1 and histone deacetylase
(DAC) 3 (Hughes et al., 2007; Yang et al., 2009; Wang et al.,
2010; reviewed by Duraisingh and Horn, 2016; reviewed by
Cestari and Stuart, 2018)(Fig. 3). The histone tri-
methyltransferase DOT1B that targets H3K76, for instance, is
required for rapid VSG-ES silencing and for an efficient transition
from an active to a silent state (Figueiredo et al., 2008). Also,
both the integrity of the NL and histone H1 are critical to
maintain condensed chromatin in silenced regions (DuBois
et al., 2012; Povelones et al., 2012). Strikingly, T. brucei lacks
H3K9me3, a well-characterized marker for heterochromatin,
and HP1 (Berriman et al., 2005), which plays a key role in var
gene silencing in Plasmodium (Brancucci et al., 2014).
Further, in Plasmodium, the histone methyltransferase SET10
colocalizes with the active var gene (Volz et al., 2012) and
NAD(+)-dependent histone deacetylases, Sir2A and Sir2B, are
required for silenci ng of different var gene subset s (Tonkin
et al., 2009), but these histone modifiers do not appear to affect
VSG silencing (Alsford et al., 2007). Indeed, in both trypano-
somes and malaria-causin g parasites, repressive heterochromatin
plays a critical role in silencing all but one antigen-coding gene
for successful antigenic variation. However, different chromatin
remodellers, histone readers/erasers and histone chaperones
appear to be involved in this process in trypanosomes and
Plasmodium (reviewed by Duraisingh and Horn, 2016).
In a broader perspective, post-transcriptional enhancement of
gene expression through spatial proximity to RNA-processing
centres might be particularly relevant in less complex eukaryotes,
where canonical transcriptional enhancers have not been identi-
fied, and particularly in Trypanosomatids, where transcriptional
regulation is limited. Nonetheless, it remains to be investigated
whether this type of regulation extends beyond VSGsinT. brucei,
and whether it plays a broader role in gene expression control in
kinetoplastids.
Notably, in T. brucei
, Hi-C and ChIP-Seq analyses revealed
that other highly transcribed loci (e.g. tandem arrays that encode
for histones, tubulin, heat shock proteins, etc.) can interact with
the SL-RNA array in the mammalian-infective stage (Faria
et al., 2020). Moreover, in insect-stage T. brucei, procyclin coding
loci also interact with the SL-RNA array (Faria et al., 2020). Given
the fact that Hi-C is a very sensitive technique, it can capture
strong and stable but also stochastic and transient interactions
(Finn and Misteli, 2019); therefore, it will be interesting to
investigate whether the interactions above are stochastic or
whether they are associated with stable and heritable structures
at the single-cell level. In other words, are there any other tran-
scription/splicing factories in T. brucei and possibly in other
related parasites? Could the SL-array act as an unconventional
and post-transcriptional enhancer? The fact that the tubulin
gene loci in T. cruzi do not colocalize with the SL-RNA arrays
(Dossin and Schenkman, 2005) does not completely rule out
this idea. This is not inconsistent with such interactions being
transient and therefore more difficult to capture my microscopy,
but could also mean that strong and stable interactions might be
restricted to specific gene families and specific developmental
stages, possibly depending on transcriptional activity. Additionally,
it is very inter esting that in T. brucei, the two SL-arra ys occupy dis-
tinct chromosome territories, essentially there are two SL-RNA
expression factories: is it because one is permanently used to sustain
the expression of the active-VSG gene?
In mammals, a high degree of heterogeneity in genome organ-
ization has been observed, suggesting that individual cells in a
population can assume many distinct, albeit related, spatial con-
formations (Finn et al., 2019). Notably, such variability does
not mean that chromatin organization has no functional rele-
vance, but rather suggests that structural heterogeneity may be
another layer impacting gene expression (Finn and Misteli,
Parasitology 1247
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
2019). In Plasmodium for instance, the 3D genome structure
appears to be strongly connected with the transcriptional activity
of specific gene families throughout the life cycle (Bunnik et al.,
2018). Whether such variability can be observed in
Trypanosomatid parasites and how that might modulate gene
expression at the single-cell level and in different developmental
stages remains to be unravelled.
Future directions
Trypanosoma brucei genome sequencing was a phenomenal turn-
ing point that marked the beginning of a new era of research.
Since then, sequencing technology, gene-editing tools, imaging
and affinity purification techniques have massively evolved, allow-
ing us to experimentally tackle long-standing questions that had
been previously untrackable.
Similarly to many pathogens, in T. brucei, the highly repetitive
nature and heterozygosity of the antigen-gene arrays had pre-
cluded a complete genome assembly. Recently, through a combin-
ation of PacBio single-molecule real-time sequencing technology
and Hi-C, the haplotype-specific assembly and scaffolding of
the long antigen-gene arrays has been successful (Muller et al.,
2018). This refined genome assembly has been proven critical
to perform further analyses on chromatin organization and
gene expression, especially regarding VSG genes. Among several
downstream analyses, which largely benefitt ed from a refined gen-
ome assembly, is Hi-C.
Hi-C and other chromosome conformational capture techni-
ques are a set of powerful molecular biology methods based on
proximity labelling, which enable the analysis of chromatin spatial
organization. These methods quantify the interaction frequency
between genomic loci that are nearby in the 3D nuclear space,
but may be far in the linear genome, allowing the identification
of enhancer–promoter contacts or chromatin loops for instance
(reviewed by Kempfer and Pombo, 2020). Hi-C studies in T. bru-
cei have identified key architectural proteins and that a specific
chromatin configuration is critical to fine-tune recombination
events; indeed, perturbation of that specific architecture triggers
switches in antigen expression (Muller et al., 2018). Further, vir-
tual 4C analyses survey the interaction frequencies between a bait
locus of interest and any other loci in the genome. In T. brucei,
such analyses have demonstrated that the active-VSG ES (but
not the silent) as well as genes encoding for other highly abundant
proteins interact with the SL-array, uncovering a potential
enhancer-like mechanism (Faria et al., 2020).
Next-generation sequencing techniques including RNA-Seq
(transcript abundance), ChIP-Seq (chromatin-association) and
CLIP-Seq (RNA-binding) have now been amply used in trypano-
somes and other Trypanosomatids. More recently, ATAC-Seq
(chromatin accessibility) and single-cell RNA-Seq have been per-
formed in T. brucei (Muller et al., 2018). The latter opens unpre-
cedented opportunities to investigate differential gene expression
during developmental transitions and inherent single-cell vari-
ability within a particular life cycle stage.
Huge improvements have been made to imaging techniques
and the fast pace of development is truly remarkable. In T. brucei,
protein and DNA loci have been recently tracked at high res o-
lution, using confocal-based or structured illumination micros-
copy (XY resolution 100–120 nm), which has been critical to
characterize specific sub-nuclear compartments (Glover et al.,
2016; Budzak et al., 2019; Faria et al., 2019, 2020). But there is
a growing need for methods that can image chromosomes with
greater genomic and optical resolution; super resolution micros-
copy can now allow an XY resolution as low as 20–30 nm. To
understand how the genome functions and regulates several key
biological processes, it is necessary to visualize many genomic
regions simultaneously, not just a few. Recently, there have been
huge breakthroughs in other systems, such as OligoFISSEQ, a
combination of three methods that employ fluorescence in situ
sequencing (FISSEQ) of barcoded Oligopaint probes to enable
the rapid visualization of multiple targeted genomic regions
(Nguyen et al., 2020). Another powerful technique is electron
cryotomography, an imaging technique used to produce high-
resolution 3D views of samples, typically biological macromole-
cules and cells. In trypanosomatids, it has been used to study fla-
gellar and mitochondrial structures but to my knowledge, not to
study supramolecular sub-nuclear complexes. For instance in
humans, it was extensively used to study the human NPC
(reviewed by Lin and Hoelz, 2019
).
The clustered regularly interspaced short palindromic repeats
(CRISPR)/CRISPR-associated system (Cas) technology has revo-
lutionized molecular biology; indeed, it is a powerful tool that
allows highly efficient and reproducible manipulation of genomic
sequences for both locus-specific and genome-wide approaches.
But its huge potential is not exclusively linked to the site-directed
nuclease activity. A catalytically inactive Cas9 (dCas9) can be used
as a universal recruitment platform in order to control transcrip-
tion, visualize DNA sequences or investigate in situ proteomes
(Anton et al., 2018; Martens et al., 2019). Indeed, for the identi-
fication of locus-associated proteins, dCas9 can be fused to a
FLAG-tag and targeted to a locus of interest; chromatin is then
crosslinked and fragmented; dCas9-bound chromatin fragments
are subsequently isolated by FLAG-specific antibodies and ana-
lysed via mass spectrometry (enChIP) (Anton et al., 2018).
Unlike enChIP, CasID requires the expression of dCas9 fused to
the promiscuous biotin ligase BirA*. After the culture medium
has been supplemented with exogenous biotin, BirA* catalyses
the addition of biotin to lysine residues of proteins that are in
close proximity to the dCas9-BirA* fusion protein. Lysis of the
cells and denaturation of proteins is then followed by affinity puri-
fication of biotinylated peptides, which are identified via tandem
mass spectrometry (Anton et al., 2018; Trinkle-Mulcahy, 2019).
Indeed, this DNA-centric system can be used to pull-down pro-
teins that associate with a specific locus; taking the VSG-ESs as
an example, this system could help identifying factors specifically
associated with the active or silent-ESs and factors involved in
gene activation or gene silencing.
Additionally, several different dCas9-based systems have been
developed to perform programmable control of spatial genome
organization, among those is the CRISPR-genome organization
(CRISPR-GO) system. It delivers a highly efficient and versatile
control over the spatial positioning of genomic loci relative to spe-
cific nuclear compartments, including the nuclear periphery, CBs
and promyelocytic leukaemia bodies to study how nuclear struc-
ture affects gene regulation and cellular function (Wang et al.,
2018). For example, in T. brucei, this could be used to bring gen-
omic loci in proximity to the SL-array or the NL and assess how
that impacts gene expression. Recently, a CasDrop system was
designed to study the formation of phase-separated compart-
ments in the nucleus by enabling liquid condensation of tran-
scriptional regulators at target loci (Shin et al., 2018). For
example, in T. brucei, this could be used to investigate the forma-
tion of VEX2 protein condensates at the active VSG-ES. CRISPR/
Cas9 technology has been successfully adapted to trypanosomes
(reviewed by Bryant et al., 2019) and proven highly versatile; it
will be interesting to see the future developments.
In summary, huge technological advances have been accom-
plished in the recent years and certainly many more will in the
near future. This burst of technological breakthroughs will hope-
fully pave the way for future discoveries on nuclear and genome
organization as well as gene expression control in African trypa-
nosomes and related organisms.
1248 Joana R. C. Faria
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
Concluding remarks
Trypanosoma brucei nuclear organization and gene expression
present several striking differences when compared to more com -
plex eukaryotes. Multiple lines of evidence strongly support that
its monogenic antigen transcription, which is critical for success-
ful antigenic variation, is enforced and facilitated by a key nuclear
architecture that involves specific inter-chromosomal interactions
and compartmentalization (possibly also modification) of specific
factors.
The molecular understanding of the mechanisms underpin-
ning gene expression control in different developmental stages
of these parasites is of great importance, as it might aid future vac-
cine and drug development efforts. For instance, acoziborole, a
single-dose oral drug to treat trypanosomiasis, was shown to tar-
get cleavage and polyadenylation specificity factor 3 (Wall et al.,
2018). Therefore, RNA process ing is now established as a clinic-
ally validated drug target in the African trypanosome.
Understanding the context within which drugs work can greatly
facilitate the drug discovery process.
Notably, recent technological advances on sequencing,
imaging and affinity purification techniques have led to important
discoveries and paved the way to novel research avenues regarding
nuclear organization and gene expression control in trypano-
somes. Indeed, we live in exciting times where the pace of tech-
nology development is phenomenal and hopefully will allow us
to address long-standing questions in infection biology that
were previously inaccessible.
Acknowledgements. I would like to thank David Horn for the many excit-
ing discussions about this subject and I would like to apologize to those whose
work was not cited because of content and length constraints.
Financial support. J.R.C.F. is a senior research associate funded by a
Wellcome Trust Investigator Award to Professor David Horn (100320/Z/12/Z).
Conflict of interest. None.
Ethical standards. Not applicable.
References
Adl SM, Bass D, Lane CE, Lukeš J, Schoch CL, Smirnov A, Agatha S, Berney
C, Brown MW, Burki F, Cárdenas P, Čepička I, Chistyakova L, Campo J,
Dunthorn M, Edvardsen B, Eglit Y, Guillou L, Hampl V, Heiss AA,
Hoppenrath M, James TY, Karnkowska A, Karpov S, Kim E, Kolisko
M, Kudryavtsev A, Lahr DJG, Lara E, Le Gall L, Lynn DH, Mann DG,
Massana R, Mitchell EAD, Morrow C, Park JS, Pawlowski JW, Powell
MJ, Richter DJ, Rueckert S, Shadwick L, Shimano S, Spiegel FW,
Torruella G, Youssef N, Zlatogursky V and Zhang Q (2019) Revisions
to the classification, nomenclature, and diversity of eukaryotes. Journal of
Eukaryotic Microbiology 66,4–119.
Alsford S and Horn D (2011) Elongator protein 3b negatively regulates ribo-
somal DNA transcription in African trypanosomes. Molecular Cellular
Biology 31, 1822–1832.
Alsford S and Horn D (2012) Cell-cycle-regulated control of VSG expression
site silencing by histones and histone chaperones ASF1A and CAF-1b in
Trypanosoma brucei. Nucleic Acids Research 40, 10150–10160.
Alsford NS, Navarro M, Jamnadass HR, Dunbar H, Ackroyd M, Murphy
NB, Gull K and Ersfeld K (2003) The identification of circular extrachro-
mosomal DNA in the nuclear genome of Trypanosoma brucei. Molecular
Microbiology 47, 277–289.
Alsford S, Kawahara T, Isamah C and Horn D (2007) A sirtuin in the
African trypanosome is involved in both DNA repair and telomeric gene
silencing but is not required for antigenic variation. Molecular
Microbiology 63, 724–736.
Alsford S, Turner DJ, Obado SO, Sanchez-Flores A, Glover L, Berriman M,
Hertz-Fowler C and Horn D (2011) High-throughput phenotyping using
parallel sequencing of RNA interference targets in the African trypano-
some. Genome Research 21, 915 –924.
Ambrósio DL, Lee JH, Panigrahi AK, Nguyen TN, Cicarelli RM and Günzl
A (2009) Spliceosomal proteomics in Trypanosoma brucei reveal new RNA
splicing factors. Eukaryotic Cell 8, 990–1000.
Ambrósio DL, Badjatia N and Günzl A (2015) The spliceosomal PRP19
complex of trypanosomes. Molecular Microbiology 95, 885–901.
Amit-Avraham I, Pozner G, Eshar S, Fastman Y, Kolevzon N, Yavin E and
Dzikowski R (2015) Antisense long noncoding RNAs regulate var gene
activation in the malaria parasite Plasmodium falciparum. Proceedings of
the National Academy of Sciences USA 112, E982–E991.
Anton T, Karg E and Bultmann S (2018) Applications of the CRISPR/Cas
system beyond gene editing. Biology Methods and Protocols 3, bpy002.
Aresta-Branco F, Sanches-Vaz M, Bento F, Rodrigues JA and Figueiredo
LM (2019) African trypanosomes expressing multiple VSGs are rapidly
eliminated by the host immune system. Proceedings of the National
Academy of Sciences USA 116, 20725–20735.
Badjatia N, Ambrósio DL, Lee JH and Günzl A (2013) Trypanosome
cdc2-related kinase 9 controls spliced leader RNA cap4 methylation and
phosphorylation of RNA polymerase II subunit RPB1. Molecular Cellular
Biology 33, 1965–1975.
Bangs JD, Crain PF, Hashizume T, McCloskey JA and Boothroyd JC (1992)
Mass spectrometry of mRNA cap 4 from trypanosomatids reveals two novel
nucleosides. Journal of Biological Chemistry 267, 9805–9815.
Barquilla A, Crespo JL and Navarro M (2008) Rapamycin inhibits trypano-
some cell growth by preventing TOR complex 2 formation. Proceedings of
the National Academy of Sciences USA 105, 14579–14584.
Belli SI (2000) Chromatin remodelling during the life cycle of trypanosoma-
tids. International Journal of Parasitology 30, 679–687.
Berriman M, et al. (2005) The genome of the African trypanosome
Trypanosoma brucei. Science (New York, N.Y.) 309, 416–422.
Brancucci NMB, Bertschi NL, Zhu L, Niederwieser I, Chin WH, Wampfler
R, Freymond C, Rottmann M, Felger I, Bozdech Z and Voss TS (2014)
Heterochromatin protein 1 secures survival and transmission of malaria
parasites. Cell Host & Microbe 16, 165–176.
Brandenburg J, Schimanski B, Nogoceke E, Nguyen TN, Padovan JC, Chait
BT, Cross GA and Günzl A (2007) Multifunctional class I transcription in
Trypanosoma brucei depends on a novel protein complex. Embo Journal 26,
4856–4866.
Briggs E, Crouch K, Lemgruber L, Lapsley C and McCulloch R (2018)
Ribonuclease H1-targeted R-loops in surface antigen gene expression sites
can direct trypanosome immune evasion. PLoS Genetics 14, e1007729.
Briggs E, Crouch K, Lemgruber L, Hamilton G, Lapsley C and McCulloch R
(2019) Trypanosoma brucei ribonuclease H2A is an essential R-loop pro-
cessing enzyme whose loss causes DNA damage during transcription initi-
ation and antigenic variation. Nucleic Acids Research 47, 9180–9197.
Bryant JM, Baumgarten S, Glover L, Hutchinson S and Rachidi N
(2019)
CRISPR in parasitology: not exactly cut and dried!. Trends in Parasitology
35, 409–422.
BudzakJ,KerryLE,AristodemouA,HallBS,WitmerK,KushwahaM,Davies
C, Pove lones ML, McDonald JR, Sur A, Myler PJ and Rudenko G (2019)
Dynamic colocaliza tion of 2 simultaneousl y ac tiv e VSG Expression sites within
a single expression-site body in Trypanosoma brucei. Proceedings of the
National Academy of Sciences of the USA 116, 16561–16570.
Bunnik EM, Cook KB, Varoquaux N, Batugedara G, Prudhomme J, Cort A,
Shi L, Andolina C, Ross LS, Brady D, Fidock DA, Nosten F, Tewari R,
Sinnis P, Ay F, Vert JP, Noble WS and Le Roch KG (2018) Changes in
genome organization of parasite-specific gene families during the
Plasmodium transmission stages. Nature Communications 9, 1910.
Capewell P, Cren-Travaillé C, Marchesi F, Johnston P, Clucas C, Benson
RA, Gorman TA, Calvo-Alvarez E, Crouzols A, Jouvion G, Jamonneau
V, Weir W, Stevenson ML, O’Neill K, Cooper A, Swar NK, Bucheton
B, Ngoyi DM, Garside P, Rotureau B and MacLeod A (2016) The skin
is a significant but overlooked anatomical reservoir for vector-borne
African trypanosomes. Elife 5, e17716.
Cestari I and Stuart K (2015) Inositol phosphate pathway controls transcrip-
tion of telomeric expression sites in trypanosomes. Proceedings of the
National Academy of Sciences of the USA 112, E2803–E2812.
Cestari I and Stuart K (2018) Transcriptional regulation of telomeric expres-
sion sites and antigenic variation in trypanosomes. Current Genomics 19,
119–132.
Cestari I, McLeland-Wieser H and Stuart K (2019) Nuclear phosphatidylino-
sitol 5-phosphatase is essential for allelic exclusion of variant surface glyco-
protein genes in trypanosomes. Molecular Cellular Biology 39,3.
Parasitology 1249
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
Chaves I, Zomerdijk J, Dirks-Mulder A, Dirks RW, Raap AK and Borst P
(1998) Subnuclear localization of the active variant surface glycoprotein
gene expression site in Trypanosoma brucei. Proceedings of the National
Academy of Sciences of the USA 95, 12328–12333.
Chaves I, Rudenko G, Dirks-Mulder A, Cross M and Borst P (1999) Control
of variant surface glycoprotein gene-expression sites in Trypanosoma bru-
cei. Embo Journal 18, 4846–4855.
Clayton C (2019) Regulation of gene expression in trypanosomatids: living
with polycistronic transcription. Open Biology 9, 190072.
Cong YS, Wright WE and Shay JW (2002) Human telomerase and its regu-
lation. Microbiology and Molecular Biology Reviews 66, 407–425, table of
contents.
Daniels JP, Gull K and Wickstead B (2010) Cell biology of the trypanosome
genome. Microbiology and Molecular Biology Reviews 74, 552–569.
Das A, Zhang Q, Palenchar JB, Chatterjee B, Cross GA and Bellofatto V
(2005) Trypanosomal TBP functions with the multisubunit transcription
factor tSNAP to direct spliced-leader RNA gene expression. Molecular
and Cellular Biology 25, 7314–7322.
Das A, Li H, Liu T and Bellofatto V (2006) Biochemical characterization of
Trypanosoma brucei RNA polymerase II. Molecular Biochemical
Parasitology 150, 201–210.
Dean S, Sunter JD and Wheeler RJ (2017) Tryptag.org: a trypanosome
genome-wide protein localisation resource. Trends in Parasitology 33,80–
82.
Dechat T, Pfleghaar K, Sengupta K, Shimi T, Shumaker DK, Solimando L
and Goldman RD (2008) Nuclear lamins: major factors in the structural
organization and function of the nucleus and chromatin. Genes and
Development 22, 832–853.
DeGrasse JA, DuBois KN, Devos D, Siegel TN, Sali A, Field MC, Rout MP
and Chait BT (2009) Evidence for a shared nuclear pore complex architec-
ture that is conserved from the last common eukaryotic ancestor. Molecular
& Cellular Proteomics 8, 2119–2130.
de Lange T (2005) Shelterin: the protein complex that shapes and safeguards
human telomeres. Genes & Development 19, 2100–2110.
Denninger V and Rudenko G (2014) FACT plays a major role in histone
dynamics affecting VSG expression site control in Trypanosoma brucei.
Molecular Microbiology 94, 945–962.
Devaux S, Lecordier L, Uzureau P, Walgraffe D, Dierick JF, Poelvoorde P,
Pays E and Vanhamme L (2006) Characterization of RNA polymerase II sub-
units of Trypanosoma brucei. Molecular Biochemical Parasitology 148,60–68.
Devaux S, Kelly S, Lecordier L, Wickstead B, Perez-Morga D, Pays E,
Vanhamme L and Gull K (2007) Diversification of function by different
isoforms of conventionally shared RNA polymerase subunits. Molecular
Biology of the Cell 18, 1293–1301.
Djikeng A, Shi H, Tschudi C and Ullu E (2001) RNA interference in
Trypanosoma brucei: cloning of small interfering RNAs provides evidence
for retroposon-derived 24–26-nucleotide RNAs. RNA 7, 1522–1530.
do Nascimento LM, Egler F, Arnold K, Papavisiliou N, Clayton C and
Erben E (2020) The trypanosome variant surface glycoprotein mRNA is
stabilized by an essential unconventional RNA-binding protein. bioRxiv,
2020.2010.2008.331769.
Dossin Fde M and Schenkman S (2005) Actively transcribing RNA polymer-
ase II concentrates on spliced leader genes in the nucleus of Trypanosoma
cruzi. Eukaryotic Cell 4, 960–970.
Dreesen O, Li B and Cross GA (2005) Telomere structure and shortening in
telomerase-deficient Trypanosoma brucei. Nucleic Acids Research 33, 4536–
4543.
DuBois KN, Alsford S, Holden JM, Buisson J, Swiderski M, Bart JM,
Ratushny AV, Wan Y, Bastin P, Barry JD, Navarro M, Horn D,
Aitchison JD, Rout MP and Field MC (2012) NUP-1 is a large coiled-coil
nucleoskeletal protein in trypanosomes with lamin-like functions. PLoS
Biology 10, e1001287.
Duraisingh MT and Horn D (2016) Epigenetic regulation of virulence gene
expression in parasitic protozoa. Cell Host & Microbe 19, 629–640.
Duraisingh MT, Voss TS, Marty AJ, Duffy MF, Good RT, Thompson JK,
Freitas-Junior LH, Scherf A, Crabb BS and Cowman AF (2005)
Heterochromatin silencing and locus repositioning linked to regulation of
virulence genes in Plasmodium falciparum. Cell 121,13–24.
Elias MC, Marques-Porto R, Freymüller E and Schenkman S (2001)
Transcription rate modulation through the Trypanosoma cruzi life cycle
occurs in parallel with changes in nuclear organisation.
Molecular
Biochemical Parasitology 112,79–90.
Engel C, Gubbey T, Neyer S, Sainsbury S, Oberthuer C, Baejen C, Bernecky
C and Cramer P (2017) Structural basis of RNA polymerase I transcription
initiation. Cell 169, 120–131.e122.
Evers R, Hammer A, Köck J, Jess W, Borst P, Mémet S and Cornelissen AW
(1989) Trypanosoma brucei contains two RNA polymerase II largest sub-
unit genes with an altered C-terminal domain. Cell 56, 585–597.
Faria J, Glover L, Hutchinson S, Boehm C, Field MC and Horn D (2019)
Monoallelic expression and epigenetic inheritance sustained by a
Trypanosoma brucei variant surface glycoprotein exclusion complex.
Nature Communications 10, 3023.
Faria J, Luzak V, Muller LSM, Brink BG, Hutchinson S, Glover L, Horn D
and Siegel TN (2020) Spatial integration of transcription and splicing in a
dedicated compartment sustains monogenic antigen expression in African
trypanosomes. Nature Microbiology, in press. doi: https://doi.org/10.1038/
s41564-020-00833-4.
Ferreira-Cerca S, Pöll G, Kühn H, Neueder A, Jakob S, Tschochner H and
Milkereit P (2007) Analysis of the in vivo assembly pathway of eukaryotic
40S ribosomal proteins. Molecular Cell 28, 446–457.
Figueiredo LM, Janzen CJ and Cross GA (2008) A histone methyltransferase
modulates antigenic variation in African trypanosomes. PLoS Biology 6,
e161.
Finn EH and Misteli T (2019) Molecular basis and biological function of
variability in spatial genome organization. Science (New York, N.Y.) 365,
eaaw9498.
Finn EH, Pegoraro G, Brandão HB, Valton AL, Oomen ME, Dekker J,
Mirny L and Misteli T (2019) Extensive heterogeneity and intrinsic vari-
ation in spatial genome organization. Cell 176, 1502–1515.e1510.
Foster HA and Bridger JM (2005) The genome and the nucleus: a marriage
made by evolution. Genome organisation and nuclear architecture.
Chromosoma 114, 212–229.
Freitas-Junior LH, Hernandez-Rivas R, Ralph SA, Montiel-Condado D,
Ruvalcaba-Salazar OK, Rojas-Meza AP, Mâncio-Silva L, Leal-Silvestre
RJ, Gontijo AM, Shorte S and Scherf A (2005) Telomeric heterochromatin
propagation and histone acetylation control mutually exclusive expression
of antigenic variation genes in malaria parasites. Cell 121,25–36.
Galganski L, Urbanek MO and Krzyzosiak WJ (2017) Nuclear speckles:
molecular organization, biological function and role in disease. Nucleic
Acids Research 45
, 10350–10368.
Gibcus JH and Dekker J (2013) The hierarchy of the 3D genome. Molecular
Cell 49, 773–782.
Glover L, Alsford S and Horn D (2013) DNA Break site at fragile subtelo-
meres determines probability and mechanism of antigenic variation in
African trypanosomes. PLoS Pathogens 9, e1003260.
Glover L, Hutchinson S, Alsford S and Horn D (2016) VEX1 Controls the
allelic exclusion required for antigenic variation in trypanosomes.
Proceedings of the National Academy of Sciences of the USA 113, 7225–7230.
Günzl A (2010) The pre-mRNA splicing machinery of trypanosomes: complex
or simplified? Eukaryotic Cell 9, 1159–1170.
Günzl A, Bruderer T, Laufer G, Schimanski B, Tu LC, Chung HM, Lee PT
and Lee MG (2003) RNA Polymerase I transcribes procyclin genes and
variant surface glycoprotein gene expression sites in Trypanosoma brucei.
Eukaryotic Cell 2, 542–551.
Guo YE, Manteiga JC, Henninger JE, Sabari BR, Dall’Agnese A, Hannett
NM, Spille JH, Afeyan LK, Zamudio AV, Shrinivas K, Abraham BJ,
Boija A, Decker TM, Rimel JK, Fant CB, Lee TI, Cisse II, Sharp PA,
Taatjes DJ and Young RA (2019) Pol II phosphorylation regulates a switch
between transcriptional and splicing condensates. Nature 572, 543–548.
Hahn S (2004) Structure and mechanism of the RNA polymerase II transcrip-
tion machinery. Nature Structural & Molecular Biology 11, 394–403.
Hashem Y, des Georges A, Fu J, Buss SN, Jossinet F, Jobe A, Zhang Q, Liao
HY, Grassucci RA, Bajaj C, Westhof E, Madison-Antenucci S and Frank
J (2013) High-resolution cryo-electron microscopy structure of the
Trypanosoma brucei ribosome. Nature 494, 385–389.
Heger P, Marin B, Bartkuhn M, Schierenberg E and Wiehe T (2012) The
chromatin insulator CTCF and the emergence of metazoan diversity.
Proceedings of the National Academy of Sciences of the USA 109, 17507–
17512.
Hernandez-Verdun D, Roussel P, Thiry M, Sirri V and Lafontaine DL
(2010) The nucleolus: structure/function relationship in RNA metabolism.
Wiley Interdisciplinary Reviews. RNA 1, 415–431.
Hertz-Fowler C, Figueiredo LM, Quail MA, Becker M, Jackson A, Bason N,
Brooks K, Churcher C, Fahkro S, Goodhead I, Heath P, Kartvelishvili M,
1250 Joana R. C. Faria
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
Mungall K, Harris D, Hauser H, Sanders M, Saunders D, Seeger K,
Sharp S, Taylor JE, Walker D, White B, Young R, Cross GA, Rudenko
G, Barry JD, Louis EJ and Berriman M (2008) Telomeric expression
sites are highly conserved in Trypanosoma brucei. PLoS ONE 3, e3527.
Hnisz D, Shrinivas K, Young RA, Chakraborty AK and Sharp PA (2017) A
phase separation model for transcriptional control. Cell 169,13–23.
Holden JM, Koreny L, Obado S, Ratushny AV, Chen WM, Chiang JH, Kelly
S, Chait BT, Aitchison JD, Rout MP and Field MC (2014) Nuclear pore
complex evolution: a trypanosome Mlp analogue functions in chromosomal
segregation but lacks transcriptional barrier activity. Molecular Biology of
the Cell 25, 1421–1436.
Hondele M, Sachdev R, Heinrich S, Wang J, Vallotton P, Fontoura BMA
and Weis K (2019) DEAD-box ATPases are global regulators of phase-
separated organelles. Nature 573, 144–148.
Hughes K, Wand M, Foulston L, Young R, Harley K, Terry S, Ersfeld K and
Rudenko G (2007) A novel ISWI is involved in VSG expression site down-
regulation in African trypanosomes. Embo Journal 26, 2400–2410.
Hury A, Goldshmidt H, Tkacz ID and Michaeli S (2009) Trypanosome
spliced-leader-associated RNA (SLA1) localization and implications for
spliced-leader RNA biogenesis. Eukaryotic Cell 8,56–68.
Isaac C, Yang Y and Meier UT (1998) Nopp140 functions as a molecular link
between the nucleolus and the coiled bodies. Journal of Cell Biology 142,
319–329.
Jehi SE, Li X, Sandhu R, Ye F, Benmerzouga I, Zhang M, Zhao Y and Li B
(2014a) Suppression of subtelomeric VSG switching by Trypanosoma brucei
TRF requires its TTAGGG repeat-binding activity. Nucleic Acids Research
42, 12899–12911.
Jehi SE, Wu F and Li B (2014b) Trypanosoma brucei TIF2 suppresses VSG
switching by maintaining subtelomere integrity. Cell Research 24, 870–885.
Karpen GH, Schaefer JE and Laird CD (1988) A Drosophila rRNA gene
located in euchromatin is active in transcription and nucleolus formation.
Genes & Development 2, 1745–1763.
Kassem A, Pays E and Vanhamme L (2014) Transcription is initiated on
silent variant surface glycoprotein expression sites despite monoallelic
expression in Trypanosoma brucei. Proceedings of the National Academy
of Sciences of the USA 111, 8943
–8948.
Kelly S, Singleton W, Wickstead B, Ersfeld K and Gull K (2006)
Characterization and differential nuclear localization of Nopp140 and a
novel Nopp140-like protein in trypanosomes. Eukaryotic Cell 5, 876–879.
Kelly S, Kramer S, Schwede A, Maini PK, Gull K and Carrington M (2012)
Genome organization is a major component of gene expression control in
response to stress and during the cell division cycle in trypanosomes. Open
Biology 2, 120033.
Kempfer R and Pombo A (2020) Methods for mapping 3D chromosome
architecture. Nature Reviews Genetics 21, 207–226.
Kieft R, Zhang Y, Marand AP, Moran JD, Bridger R, Wells L, Schmitz RJ
and Sabatini R (2020) Identification of a novel base J binding protein com-
plex involved in RNA polymerase II transcription termination in trypano-
somes. PLoS Genetics 16, e1008390.
Kim J, Venkata NC, Hernandez Gonzalez GA, Khanna N and Belmont AS
(2020) Gene expression amplification by nuclear speckle association.
Journal of Cell Biology 219,1.
Kornblihtt AR, de la Mata M, Fededa JP, Munoz MJ and Nogues G (2004)
Multiple links between transcription and splicing. RNA 10, 1489–1498.
Koumandou VL, Natesan SK, Sergeenko T and Field MC (2008) The tryp-
anosome transcriptome is remodelled during differentiation but displays
limited responsiveness within life stages. BMC Genomics 9, 298.
Landeira D and Navarro M (2007) Nuclear repositioning of the VSG
Promoter during developmental silencing in Trypanosoma brucei. The
Journal of Cell Biology 176, 133–139.
Landeira D, Bart JM, Van Tyne D and Navarro M (2009) Cohesin regulates
VSG monoallelic expression in trypanosomes. The Journal of Cell Biology
186, 243–254.
Lee JH, Nguyen TN, Schimanski B and Günzl A (2007) Spliced leader RNA
gene transcription in Trypanosoma brucei requires transcription factor
TFIIH. Eukaryotic Cell 6, 641–649.
Lee JH, Cai G, Panigrahi AK, Dunham-Ems S, Nguyen TN, Radolf JD,
Asturias FJ and Günzl A (2010) A TFIIH-associated mediator head is a
basal factor of small nuclear spliced leader RNA gene transcription in early-
diverged trypanosomes. Molecular and Cellular Biology 30, 5502–5513.
Lemieux JE, Kyes SA, Otto TD, Feller AI, Eastman RT, Pinches RA,
Berriman M, Su XZ and Newbold CI (2013) Genome-wide profiling of
chromosome interactions in
Plasmodium falciparum characterizes nuclear
architecture and reconfigurations associated with antigenic variation.
Molecular Microbiology 90, 519–537.
Liang XH, Liu Q, Liu L, Tschudi C and Michaeli S (2006) Analysis of spli-
ceosomal complexes in Trypanosoma brucei and silencing of two splicing
factors Prp31 and Prp43. Molecular Biochemical Parasitology 145,29–39.
Lin DH and Hoelz A (2019) The structure of the nuclear pore complex (an
update). Annual Review of Biochemistry 88, 725–783.
Liu Z, Gutierrez-Vargas C, Wei J, Grassucci RA, Ramesh M, Espina N, Sun
M, Tutuncuoglu B, Madison-Antenucci S, Woolford JL Jr, Tong L and
Frank J (2016) Structure and assembly model for the Trypanosoma cruzi
60S ribosomal subunit. Proceedings of the National Academy of Sciences
of the USA 113, 12174–12179.
López-Farfán D, Bart JM, Rojas-Barros DI and Navarro M (2014)
SUMOylation by the E3 ligase TbSIZ1/PIAS1 positively regulates VSG
expression in Trypanosoma brucei. PLoS Pathogens 10, e1004545.
Machyna M, Neugebauer KM and StaněkD(2015) Coilin: the first 25 years.
RNA Biology 12, 590–596.
Maishman L, Obado SO, Alsford S, Bart JM, Chen WM, Ratushny AV,
Navarro M, Horn D, Aitchison JD, Chait BT, Rout MP and Field MC
(2016) Co-dependence between trypanosome nuclear lamina components
in nuclear stability and control of gene expression. Nucleic Acids Research
44, 10554–10570.
Martens KJA, van Beljouw SPB, van der Els S, Vink JNA, Baas S, Vogelaar
GA, Brouns SJJ, van Baarlen P, Kleerebezem M and Hohlbein J (2019)
Visualisation of dCas9 target search in vivo using an open-microscopy
framework. Nature Communications 10, 3552.
Martínez-Calvillo S, Florencio-Martínez LE and Nepomuceno-Mejía T
(2019) Nucleolar structure and function in trypanosomatid protozoa.
Cells 8, 421.
Merkenschlager M and Odom DT (2013) CTCF and cohesin: linking gene
regulatory elements with their targets. Cell 152, 1285–1297.
Michaeli S (2011) Trans-splicing in trypanosomes: machinery and its impact
on the parasite transcriptome. Future Microbiology 6, 459–474.
Millau JF and Gaudreau L (2011) CTCF, cohesin, and histone variants: con-
necting the genome. Biochemical Cellular Biology 89, 505
–513.
Misteli T (2007) Beyond the sequence: cellular organization of genome func-
tion. Cell 128, 787–800.
Monahan K and Lomvardas S (2015) Monoallelic expression of olfactory
receptors. Annual Review of Cell and Developmental Biology 31, 721–740.
Monahan K, Horta A and Lomvardas S (2019) LHX2- and LDB1-mediated
trans interactions regulate olfactory receptor choice. Nature 565, 448–453.
Muller LSM, Cosentino RO, Forstner KU, Guizetti J, Wedel C, Kaplan N,
Janzen CJ, Arampatzi P, Vogel J, Steinbiss S, Otto TD, Saliba AE,
Sebra RP and Siegel TN (2018) Genome organization and DNA accessibil-
ity control antigenic variation in trypanosomes. Nature 563, 121–125.
Nanavaty V, Sandhu R, Jehi SE, Pandya UM and Li B (2017) Trypanosoma
brucei RAP1 maintains telomere and subtelomere integrity by suppressing
TERRA and telomeric RNA:DNA hybrids. Nucleic Acids Research 45,
5785–5796.
Narayanan MS and Rudenko G (2013) TDP1 Is an HMG chromatin protein
facilitating RNA polymerase I transcription in African trypanosomes.
Nucleic Acids Research 41, 2981–2992.
Navarro M and Cross GA (1996) DNA rearrangements associated with mul-
tiple consecutive directed antigenic switches in Trypanosoma brucei.
Molecular Cellular Biology 16, 3615–3625.
Navarro M and Gull K (2001) A pol I transcriptional body associated with
VSG mono-allelic expression in Trypanosoma brucei. Nature 414, 759–763.
Navarro M, Peñate X and Landeira D (2007) Nuclear architecture underlying
gene expression in Trypanosoma brucei. Trends in Microbiology 15, 263–270.
Nepomuceno-Mejía T, Lara-Martínez R, Cevallos AM, López-Villaseñor I,
Jiménez-García LF and Hernández R (2010) The Trypanosoma cruzi
nucleolus: a morphometrical analysis of cultured epimastigotes in the expo-
nential and stationary phases. FEMS Microbiology Letters 313,41–46.
Nguyen TN, Schimanski B, Zahn A, Klumpp B and Günzl A (2006)
Purification of an eight subunit RNA polymerase I complex in
Trypanosoma brucei.
Molecular Biochemical Parasitology 149,27–37.
Nguyen TN, Schimanski B and Günzl A (2007) Active RNA polymerase I of
Trypanosoma brucei harbors a novel subunit essential for transcription.
Molecular Cellular Biology 27, 6254–6263.
Nguyen TN, Nguyen BN, Lee JH, Panigrahi AK and Günzl A (2012)
Characterization of a novel class I transcription factor A (CITFA) subunit
Parasitology 1251
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
that is indispensable for transcription by the multifunctional RNA polymer-
ase I of Trypanosoma brucei. Eukaryotic Cell 11, 1573–1581.
Nguyen HQ, Chattoraj S, Castillo D, Nguyen SC, Nir G, Lioutas A,
Hershberg EA, Martins NMC, Reginato PL, Hannan M, Beliveau BJ,
Church GM, Daugharthy ER, Marti-Renom MA and Wu CT (2020)
3D Mapping and accelerated super-resolution imaging of the human gen-
ome using in situ sequencing. Nature Methods 17, 822–832.
Obado SO, Brillantes M, Uryu K, Zhang W, Ketaren NE, Chait BT, Field
MC and Rout MP (2016) Interactome mapping reveals the evolutionary
history of the nuclear pore complex. PLoS Biology 14, e1002365.
Ogbadoyi E, Ersfeld K, Robinson D, Sherwin T and Gull K (2000)
Architecture of the Trypanosoma brucei nucleus during interphase and
mitosis. Chromosoma 108, 501–513.
Ori A, Banterle N, Iskar M, Andrés-Pons A, Escher C, Khanh Bui H,
Sparks L, Sol is-Mezarino V, Rinner O, Bork P, Lemke EA and Beck
M (20 13) Cell type-specific nuc lear pores: a case in point for context-
dependent stoichiometry of molecular machines. Molecular Systems
Biology 9,648.
Ottaviani A, Gilson E and Magdinier F (2008) Telomeric position effect:
from the yeast paradigm to human pathologies? Biochimie 90,93–107.
Otto TD, Böhme U, Sanders M, Reid A, Bruske EI, Duffy CW, Bull PC,
Pearson RD, Abdi A, Dimonte S, Stewart LB, Campino S, Kekre M,
Hamilton WL, Claessens A, Volkman SK, Ndiaye D, Amambua-Ngwa
A, Diakite M, Fairhurst RM, Conway DJ, Franck M, Newbold CI and
Berriman M (2018) Long read assemblies of geographically dispersed
Plasmodium falciparum isolates reveal highly structured subtelomeres.
Wellcome Open Research 3, 52.
Outters P, Jaeger S, Zaarour N and Ferrier P (2015) Long-range control of V
(D)J recombination & allelic exclusion: modeling views. Advances in
Immunology 128, 363–413.
Palfi Z, Lücke S, Lahm HW, Lane WS, Kruft V, Bragado-Nilsson E,
Séraphin B and Bindereif A (2000) The spliceosomal snRNP core complex
of Trypanosoma brucei: cloning and functional analysis reveals seven Sm
protein constituents. Proceedings of the National Academy of Sciences of
the USA 97, 8967–8972.
Povelones ML, Gluenz E, Dembek M, Gull K and Rudenko G (2012) Histone
H1 plays a role in heterochromatin formation and VSG expression site
silencing in Trypanosoma brucei. PLoS Pathogens 8, e1003010.
Preusser C, Palfi Z and Bindereif A (2009) Special Sm core complex func-
tions in assembly of the U2 small nuclear ribonucleoprotein of
Trypanosoma brucei. Eukaryotic Cell 8, 1228–1234.
Prohaska K and Williams N (2009) Assembly of the Trypanosoma brucei 60S
ribosomal subunit nuclear export complex requires trypanosome-specific
proteins P34 and P37. Eukaryotic Cell 8,77–87.
Proudfoot NJ, Furger A and Dye MJ (2002) Integrating mRNA processing
with transcription. Cell 108, 501–512.
Quinodoz SA, Ollikainen N, Tabak B, Palla A, Schmidt JM, Detmar E, Lai
MM, Shishkin AA, Bhat P, Takei Y, Trinh V, Aznauryan E, Russell P,
Cheng C, Jovanovic M, Chow A, Cai L, McDonel P, Garber M and
Guttman M (2018) Higher-order inter-chromosomal hubs shape 3D gen-
ome organization in the nucleus. Cell 174, 744–757.e724.
Rao SS, Huntley MH, Durand NC, Stamenova EK, Bochkov ID, Robinson
JT, Sanborn AL, Machol I, Omer AD, Lander ES and Aiden EL (2014) A
3D map of the human genome at kilobase resolution reveals principles of
chromatin looping. Cell 159, 1665–1680.
Raska I, Shaw PJ and Cmarko D (2006) Structure and function of the nucle-
olus in the spotlight. Current Opinion in Cell Biology 18, 325–334.
Razin SV and Gavrilov AA (2020) The role of liquid-liquid phase separation
in the compartmentalization of cell nucleus and spatial genome organiza-
tion. Biochemistry (Mosc) 85, 643–650.
Reis H, Schwebs M, Dietz S, Janzen CJ and Butter F (2018) TelAP1 links
telomere complexes with developmental expression site silencing in
African trypanosomes. Nucleic Acids Research 46, 2820–2833.
Reynolds DL, Hofmeister BT, Cliffe L, Siegel TN, Anderson BA, Beverley
SM, Schmitz RJ and Sabatini R (2016) Base J represses genes at the end
of polycistronic gene clusters in Leishmania major by promoting RNAP
II termination. Molecular Microbiology 101, 559–574.
Rico E, Jeacock L, Kovářová J and Horn D (2018) Inducible high-efficiency
CRISPR-Cas9-targeted gene editing and precision base editing in African
trypanosomes. Scientific Reports 8, 7960.
RoutMP,ObadoSO,SchenkmanSandFieldMC(2017) Specialising the parasite
nucleus: pores, lamins, chroma tin, and diversity. PLoS Pathogens 13, e1006170.
Rubbi CP and Milner J (2003) Disruption of the nucleolus mediates stabiliza-
tion of p53 in response to DNA damage and other stresses. Embo Journal
22, 6068–6077.
Saha A, Nanavaty VP and Li B (2020) Telomere and subtelomere R-loops and
antigenic variation in trypanosomes. Journal of Molecular Biology 432,
4167–4185.
Sandhu R, Sanford S, Basu S, Park M, Pandya UM, Li B and Chakrabarti K
(2013) A trans-spliced telomerase RNA dictates telomere synthesis in
Trypanosoma brucei. Cell Research 23, 537–551.
Sawyer IA, Sturgill D, Sung MH, Hager GL and Dundr M (2016) Cajal body
function in genome organization and transcriptome diversity. Bioessays 38,
1197–1208.
Schimanski B, Nguyen TN and Günzl A (2005) Characterization of a multi-
subunit transcription factor complex essential for spliced-leader RNA gene
transcription in Trypanosoma brucei. Molecular Cell Biology 25, 7303–7313.
Schimanski B, Brandenburg J, Nguyen TN, Caimano MJ and Günzl A
(2006) A TFIIB-like protein is indispensable for spliced leader RNA gene
transcription in Trypanosoma brucei. Nucleic Acids Research 34, 1676–1684.
Schoenfelder S and Fraser P (2019) Long-range enhancer-promoter contacts
in gene expression control. Nature Reviews Genetics 20, 437–455.
Schulz D, Mugnier MR, Paulsen EM, Kim HS, Chung CW, Tough DF,
Rioja I, Prinjha RK, Papavasiliou FN and Debler EW (2015)
Bromodomain proteins contribute to maintenance of bloodstream form
stage identity in the African trypanosome. PLoS Biology 13, e1002316.
Schulz D, Zaringhalam M, Papavasiliou FN and Kim HS (2016) Base J and
H3.V regulate transcriptional termination in Trypanosoma brucei. PLoS
Genetics 12, e1005762.
Shevelyov YY and Ulianov SV (2019) The nuclear lamina as an organizer of
chromosome architecture. Cells 8, 136.
Shin Y, Chang YC, Lee DSW, Berry J, Sanders DW, Ronceray P, Wingreen
NS, Haataja M and Brangwynne CP (2018) Liquid nuclear condensates
mechanically sense and restructure the genome. Cell 175, 1481–1491.e1413.
Shpargel KB, Ospina JK, Tucker KE, Matera AG and Hebert MD (2003)
Control of Cajal body number is mediated by the coilin C-terminus.
Journal of Cell Science 116, 303–312.
Spann TP, Moir RD, Goldman AE, Stick R and Goldman RD
(1997)
Disruption of nuclear lamin organization alters the distribution of replication
factors and inhibits DNA synthesis. Journal of Cell Biology 136, 1201–1212.
Srivastava A, Badjatia N, Lee JH, Hao B and Günzl A (2018) An RNA poly-
merase II-associated TFIIF-like complex is indispensable for SL RNA gene
transcription in Trypanosoma brucei. Nucleic Acids Research 46, 1695–1709.
Strom AR and Brangwynne CP (2019) The liquid nucleome – phase transi-
tions in the nucleus at a glance. Journal of Cell Science 132, jcs235093.
Tan J and Lan L (2020) The DNA secondary structures at telomeres and gen-
ome instability. Cell & Bioscience 10, 47.
Tkacz ID, Lustig Y, Stern MZ, Biton M, Salmon-Divon M, Das A,
Bellofatto V and Michaeli S (2007) Identification of novel
snRNA-specific Sm proteins that bind selectively to U2 and U4 snRNAs
in Trypanosoma brucei. RNA 13,30–43.
Tkacz ID, Gupta SK, Volkov V, Romano M, Haham T, Tulinski P,
Lebenthal I and Michaeli S (2010) Analysis of spliceosomal proteins in
Trypanosomatids reveals novel functions in mRNA processing. Journal of
Biological Chemistry 285, 27982–27999.
Tonkin CJ, Carret CK, Duraisingh MT, Voss TS, Ralph SA, Hommel M,
Duffy MF, Silva LM, Scherf A, Ivens A, Speed TP, Beeson JG and
Cowman AF (2009) Sir2 paralogues cooperate to regulate virulence genes
and antigenic variation in Plasmodium falciparum. PLoS Biology 7, e84.
Trindade S, Rijo-Ferreira F, Carvalho T, Pinto-Neves D, Guegan F,
Aresta-Branco F, Bento F, Young SA, Pinto A, Van Den Abbeele J,
Ribeiro RM, Dias S, Smith TK and Figueiredo LM (2016)
Trypanosoma brucei parasites occupy and functionally adapt to the adipose
tissue in mice. Cell Host & Microbe 19, 837–848.
Trinkle-Mulcahy L (2019) Recent advances in proximity-based labeling meth-
ods for interactome mapping. F1000Research 8.
Uzureau P, Daniels JP, Walgraffe D, Wickstead B, Pays E, Gull K and
Vanhamme L (2008) Identification and characterization of two trypano-
some TFIIS proteins exhibiting particular domain architectures and differ-
ential nuclear localizations. Molecular Microbiology 69, 1121–1136.
Vanhamme L, Poelvoorde P, Pays A, Tebabi P, Van Xong H and Pays E
(2000) Differential RNA elongation controls the variant surface glycopro-
tein gene expression sites of Trypanosoma brucei. Molecular Microbiology
36, 328–340.
1252 Joana R. C. Faria
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
Viegas IJ, de Macedo JP, De Niz M, Rodrigues JA, Aresta-Branco F, Jaffrey
SR and Figueiredo LM (2020) N6-methyladenosine in poly(A) tails stabil-
ize VSG transcripts. bioRxiv. doi: 10.1101/2020.01.30.925776.
Volz JC, Bártfai R, Petter M, Langer C, Josling GA, Tsuboi T , Schw ach F , Baum
J, Rayner JC, Stunnenberg HG, Duffy MF and Cowman AF (2012) PfSET10,
a Plasmodium falciparum methyltransfer ase, maintains the activ e var gene in a
poised state during parasite division. Cell Host & Microbe 11,7–18.
Walgraffe D, Devaux S, Lecordier L, Dierick JF, Dieu M, Van den Abbeele
J, Pays E and Vanhamme L (2005) Characterization of subunits of the
RNA polymerase I complex in Trypanosoma brucei. Molecular
Biochemical Parasitology 139, 249–260.
Wall RJ, Rico E, Lukac I, Zuccotto F, Elg S, Gilbert IH, Freund Y, Alley
MRK, Field MC, Wyllie S and Horn D (2018) Clinical and veterinary try-
panocidal benzoxaboroles target CPSF3. Proceedings of the National
Academy of Sciences of the USA 115, 9616–9621.
Wang QP, Kawahara T and Horn D (2010) Histone deacetylases play distinct
roles in telomeric VSG expression site silencing in African trypanosomes.
Molecular Microbiology 77, 1237–1245.
Wang H, Xu X, Nguyen CM, Liu Y, Gao Y, Lin X, Daley T, Kipniss NH, La
Russa M and Qi LS (2018) CRISPR-mediated programmable 3D genome
positioning and nuclear organization. Cell 175, 1405–1417.e1414.
Wedel C, Förstner KU, Derr R and Siegel TN (2017) GT-rich promoters can
drive RNA pol II transcription and deposition of H2A.Z in African trypa-
nosomes. Embo Journal 36, 2581–2594.
Wickstead B, Ersfeld K and Gull K (2004) The small chromosomes of
Trypanosoma brucei involved in antigenic variation are constructed around
repetitive palindromes. Genome Research 14, 1014–1024.
Yang X, Figueiredo LM, Espinal A, Okubo E and Li B (2009) RAP1 is essen-
tial for silencing telomeric variant surface glycoprotein genes in
Trypanosoma brucei. Cell 137,99–109.
Parasitology 1253
https://doi.org/10.1017/S0031182020002437 Published online by Cambridge University Press
