
1
DNA Wrap: Packaging Matters
An Introduction to Epigenetics
UNC-Chapel Hill’s Superfund Research Program
What causes the physical appearance and health status of identical twins to diverge with age? In this lesson,
students learn that the environment can alter the way our genes are expressed, making even identical twins
different. After watching a PBS video, A Tale of Two Mice, and reviewing data presented in the Environmental
Health Perspectives article Maternal Genistein Alters Coat Color and Protects Avy Mouse Offspring from Obesity by Modifying
the Fetal Epigenome, students learn about epigenetics and its role in regulating gene expression.
Author
Dana Haine, MS
University of North Carolina at Chapel Hill Superfund Research Program
Reviewers
Dana Dolinoy, PhD
University of Michigan
Rebecca Fry, PhD
University of North Carolina at Chapel Hill Superfund Research Program
Banalata Sen, PhD, Audrey Pinto, PhD, Susan Booker, Dorothy Ritter
Environmental Health Perspectives
The funding for development of this lesson was provided by the National Institute of Environmental Health
Sciences and the UNC Superfund Program.
Learning Objectives
By the end of this lesson students should be able to:
define the term “epigenetics”
describe DNA methylation as a mechanism for inhibiting gene transcription
describe how gene expression can vary among genetically identical offspring
Alignment to NC Essential Science Standards for Biology
Bio.3.1 Explain how traits are determined by the structure and function of DNA.
Bio.3.2.3 Explain how the environment can influence the expression of genetic traits.
Bio.4.1 Understand how biological molecules are essential to the survival of living organisms.
Bio.4.1.2 Summarize the relationship among DNA, proteins and amino acids in carrying out the work
of cells and how this is similar in all organisms.
Bio.3.3 Understand the application of DNA technology.
Bio.3.3.1 Interpret how DNA is used for comparison and identification of organisms.
Note to Teacher: This lesson is best conducted after students have learned about general DNA structure and
function; transcription and translation; general regulation of gene expression.
2
Skills Used or Developed
Communication (note-taking, oral, written – including summarization)
Comprehension (listening, reading)
Critical thinking and response
Graph reading
Figures (reading legends)
Class Time Required
45-60 minutes
Depending on student proficiency level, this lesson can be completed as a homework assignment to encourage
independent student work and/or to save class time.
Materials
Computer with Internet access and audio (sound) capabilities
LCD Projector
Accompanying Powerpoint slide library (optional)
Student worksheet, one copy per student
Teacher Preparation
Ensure students have a basic understanding of DNA structure and function prior to introducing the
concept of epigenetics. Students should already be familiar with the following terminology:
o Chromatin
o Chromosome
o Deoxyribonucleic acid (DNA)
o Gene
o Gene expression/regulation
o Histone proteins
o Nitrogenous base
o Nucleoprotein
o Nucleotides
o Phosphodiester bond
o Promoter
o Ribonucleic acid (RNA)
o Transcription
o Transcription factors
o Translation
Review the Background Information, Procedure and Student Instructions for this activity.
Additional resources are listed in the Resources section.
Make copies of the Student Worksheet, one per student.
Background Information
Deoxyribonucleic acid (DNA) is a large, complex molecule (macromolecule) that contains the genetic
code or the information needed to direct the activities of a cell and for transmission of this information to
the next generation. A single DNA strand is made up of building blocks called nucleotides that are
connected together like a chain. Each DNA nucleotide is composed of a nitrogenous base—either
adenosine (A), guanine (G), thymine (T), or cytosine (C)—a five-carbon deoxyribose sugar (S), and a
phosphate group (P). A gene is a specific sequence of nucleotides within a DNA strand that provides the
instructions necessary to carry out a particular activity.
DNA exists as a double-stranded polymer of nucleotides that forms a helix in which two DNA strands run
anti-parallel to one another and interact via hydrogen bonds between the nitrogenous bases. The
hydrogen bonds between the nitrogenous bases can be broken to allow the DNA strands to separate during
3
DNA replication and gene expression, which occurs when the nucleotide sequence of a gene is copied into
ribonucleic acid (RNA) during a process called transcription. For genes that encode proteins, DNA is
copied into messenger RNA (mRNA), which then directs the synthesis of proteins during the process
known as translation. Gene expression is highly regulated in order to control a cell’s activities; thus, the
timing and amount of RNA and protein generated from a given gene varies depending on the cell’s
activities. Disruption of gene expression regulation leads to diseases such as cancer.
Inside cells, DNA is packaged around proteins called histones; this DNA–protein (nucleoprotein)
complex is called chromatin. Histones act like “spools” around which DNA is wrapped. In humans, each
cell contains approximately 2 meters of DNA; however, because of the wrapping of DNA around histones,
the condensed DNA is approximately 120 micrometers long!
This DNA “packaging” in the form of chromatin plays a key role in the regulation of gene expression. The
nucleoprotein inside cells serves as a docking site for the different proteins and enzymes and their
interactions required for DNA replication, transcription, recombination, and repair.
This lesson introduces students to the emerging field of epigenetics. Epigenetics literally means “on top
of or in addition to genetics,” and is the study of changes in gene expression not accompanied by alterations
in DNA sequence. In parallel to the term genome, which defines the complete set of genetic information
contained in the DNA of an organism, epigenome refers to the complete set of epigenetic pathways in an
organism. Epigenetic modifications to DNA exert profound influences on gene activity. For example,
studies suggest that epigenetic variation may be responsible for subtle differences in appearance and
behavior of identical twins. Identical twins are more epigenetically similar early in life but show remarkable
divergence with age.
Epigenetic pathways such as DNA methylation and histone modifications interact with each other to
regulate expression of genes. One of the most common and well-characterized epigenetic pathways is DNA
methylation. DNA methylation occurs when an enzyme called a methyltransferase covalently attaches a
methyl (-CH3) group to a cytosine base that is adjacent to a guanine base
(see Figure 1). Such sites where a cytosine is adjacent to guanine via a
phosphodiester bond are called CpG sites. Scientists have observed that
DNA methylation occurs predominately along places on the DNA
strand that are rich in CpG pairs. One type of CpG-rich region is a CpG
island. CpG islands are associated with approximately 60-70% of
mammalian genes, and most CpG islands are unmethylated in normal
mammalian cells. Thus, changes in methylation patterns at CpG islands
can interfere with normal gene expression by altering the transcriptional
competency of a gene’s promoter. Genes that are essential for a cell’s function are not methylated. In
contrast, inactive genes are usually methylated to suppress their expression.
While DNA methylation is involved in normal control of gene expression, changes in the extent of DNA
methylation can contribute to cancer or disease by silencing genes that that should otherwise be active or
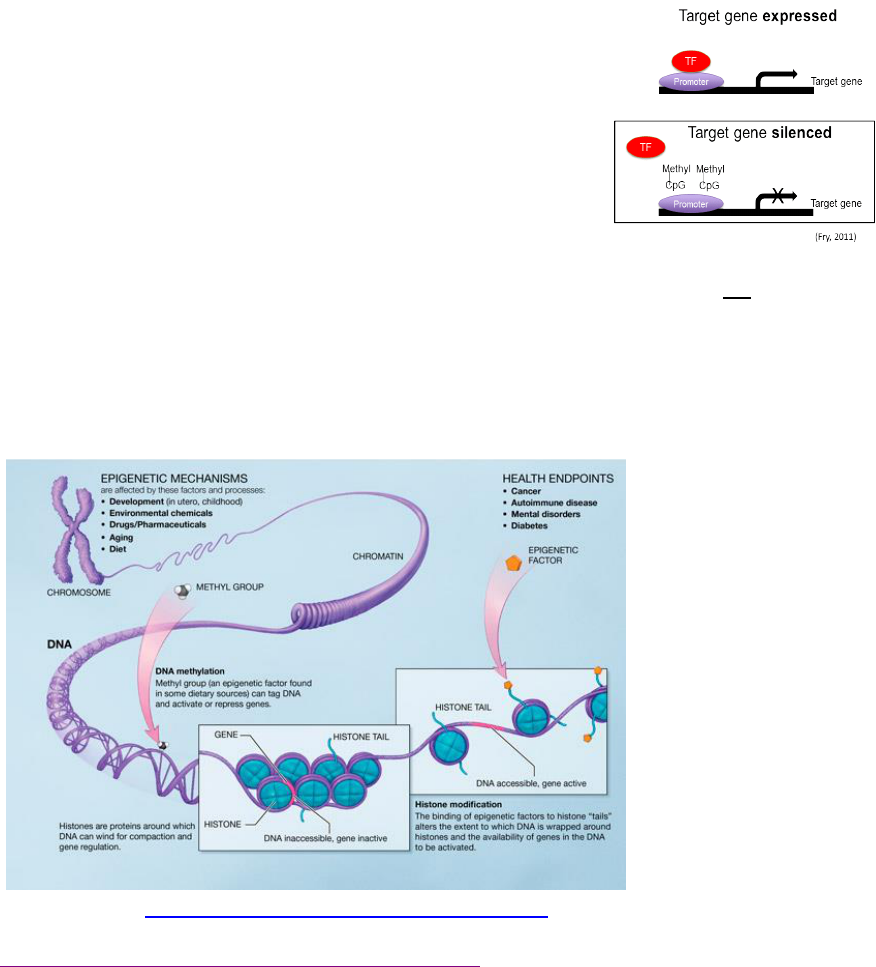
expressed or by causing expression of genes that are usually inactive. Methylation is one mechanism for
suppressing (or silencing) gene transcription by preventing one or more transcription factors (TF) and
thus RNA polymerase from accessing a gene’s promoter which is required for transcribing DNA into
RNA (see Figure 2).
Figure 1: This representative DNA helix
depicts two methyl groups (M)
covalently attached to two cyotosine
bases in DNA.
C G T A C A C G A C A C G A T
G C A T G T G C T G T G C T A
4
The histone proteins that hold DNA tightly wound inside each cell can also be
modified by methylation or other modifications such as acetylation or
phosphorylation. When too much or too little of a given histone modification occurs,
it affects a gene’s expression and consequently its function, which causes unwanted
alterations in the cell, potentially resulting in disease.
Epigenetic modifications can be maintained and inherited by daughter cells during
mitosis and to a lesser extent during meiosis. Therefore, epigenetic modifications that
occur in utero can be passed on to subsequent generations. Environmental factors such
as exposure to heavy metals (arsenic, nickel) and cigarette smoke, and dietary factors
such as vitamin and folate deficiencies have been linked to abnormal changes in
epigenetic pathways, suggesting that an individual’s environment plays an important
role in shaping their epigenome. Epigenetic changes have been observed in different
stages of cancer progression, in the process of aging, and in other human diseases such
as Alzheimer’s disease, diabetes and obesity.
Source:
https://commonfund.nih.gov/epigenomics/figure
In A Tale of Two Mice (http://www.pbs.org/wgbh/nova/genes/mice.html), the narrator discusses the difference
in coat color between two genetically identical mice. The obese, yellow mouse has an unmethylated Agouti gene,
which is constantly being expressed (when it normally should be “off ” or silenced), while
her sister, the brown mouse, has a methylated Agouti gene that has permanently been turned “off ” and thus is
not expressed. Although genetically identical in terms of the DNA sequences they’ve inherited from their
mother and father (the mice are inbred), epigenetic modifications have led one mouse to be overweight and
more susceptible to diabetes and cancer. This difference in gene expression between the genetically identical
mice can be attributed to differential gene expression as a result of epigenetic modifications.
Figure 2: The addition of methyl
groups to CpG islands common to
promoters is one mechanism for
suppressing (or silencing) gene
transcription (Image: Fry, 2011).

Procedure
1. As an engagement activity, go to http://www.pbs.org/wgbh/nova/body/epigenetic-mice.html and launch
the interactive slide show titled A Tale of Two Mice. Show students Chapter 1: The Agouti Sisters (50 seconds)
to generate student interest.
2. Distribute copies of the Student Worksheet and ask students to work in pairs to complete Step 1.
3. Promote a brief class discussion by asking students to share their answers to Step 1, questions 1 and 2, with
the class.
What does it mean to say that two individuals are genetically identical?
How can two genetically identical mice look so different?
4. Tell students that the answer to these questions lies in how DNA is packaged inside cells and then invite
students to complete Step 2 on their worksheet. The amount of time students need for this step depends
on the extent to which DNA and chromosome structure has already been covered in class.
5. Review student answers to Step 2 before proceeding to Step 3.
6. Next, introduce students to the concept that changes in gene expression can occur without changes in the
DNA sequence of genes (mutation). Describe the process of DNA methylation as a means of silencing
transcription of a gene (as described in the Background section).
7. To conclude this description of DNA methylation, return to the audio slide show A Tale of Two Mice. and
show students Chapter 2: The Epigenome (1:06) and at least the first 15 seconds of the next chapter,
Switching on the Agouti Gene – which explains that in the yellow, obese mouse, the agouti gene is
unmetlyated and turned on all of the time while in the brown mouse the gene is completely methylated
and shut down.
8. Ask students to complete Step 3 by summarizing in their own words how DNA methylation affects DNA
structure and function. Review student responses as a class before proceeding.
9. Next, ask students to read Step 4, examine Figures 4 and 5 from the featured Environmental Health Perspectives
article and answer the corresponding questions. Optional: Expose students to the original article and have them read
and discuss all or part of this scientific publication.
10. Review student responses as a class before allowing them to proceed to Step 5.
11. Ask students to read Step 5, and with a partner, discuss how the authors’ conclusions are significant to
them.
12. Discuss student responses as a class to conclude this lesson.
13. Optional: If time permits, you may choose to finish showing A Tale of Two Mice or the entire NOVA special
Epigenetics, available at: http://www.pbs.org/wgbh/nova/body/epigenetics.html
Assessment Options
Ask students to turn in their completed worksheets (Answer Key provided).
Ask student to summarize, in their words, what they learned during this activity.
Ask students to construct a concept map using critical vocabulary terms (see below) along with
vocabulary terms from page 2 to demonstrate they understand the concept of epigenetics in the
context of DNA structure and function.
Critical Vocabulary
Definitions for the terms below were obtained from the National Institute of Health, Genetics Home Reference Handbook
http://ghr.nlm.nih.gov/handbook/howgeneswork/epigenome
Epigenetics: refers to heritable changes in the regulation of the expression of gene activity without alteration
of genetic structure.

Epigenetic modifications: chemical compounds (e.g. methyl groups) that are added to single genes can
regulate their activity; these modifications are known as epigenetic changes. These chemical “tags” can remain
in place as cells divide and in some cases can be inherited through generations.
Epigenome: refers to all of the chemical compounds that have been added to the entirety of one’s DNA
(genome) as a way to regulate the activity (expression) of all the genes within the genome. Environmental
influences, such as a person’s diet and exposure to pollutants, can also impact the epigenome.
Methylation: small molecules called methyl groups, each consisting of one carbon atom and three hydrogen
atoms, are covalently attached to segments of DNA by an enzyme known as a methyltransferase. When methyl
groups are added to a particular gene, that gene is turned off or silenced, and no protein is produced from that
gene.
Methyltransferase: refers the enzyme that covalently attaches a methyl group to DNA.
Resources
Bob Weinhold. Epigenetics: The Science of Change. Environmental Health Perspectives. 2006.
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1392256/
National Human Genome Research Institute, Epigenomics Fact Sheet.
http://www.genome.gov/27532724
National Institute of Health, Genetics Home Reference, What is the Epigenome?
http://ghr.nlm.nih.gov/handbook/howgeneswork/epigenome
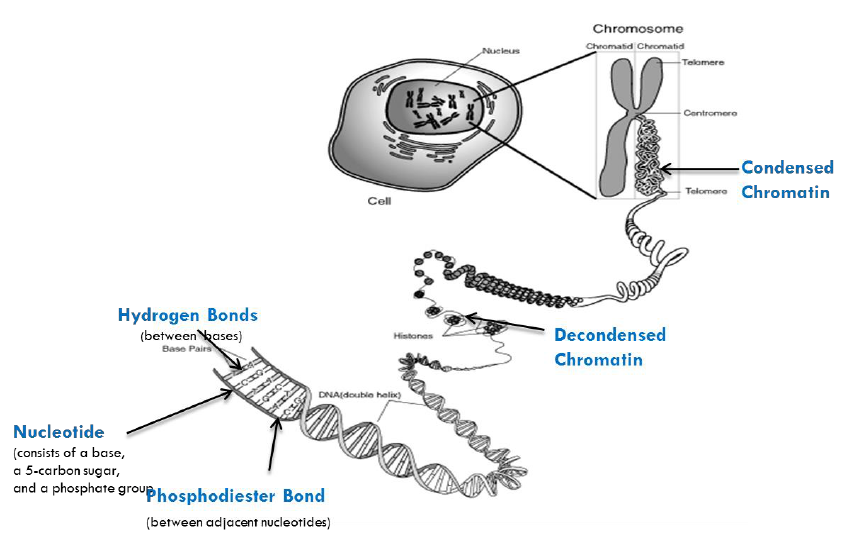
National Institute of Health, A Scientific Illustration of How Epigenetic Mechanisms Can Affect Health
https://commonfund.nih.gov/epigenomics/figure
NOVA, Ghost in Your Genes.
http://www.pbs.org/wgbh/nova/genes/
Scitable by Nature Education, Epigentic Influences and Disease. http://www.nature.com/scitable/topicpage/Epigenetic-
Influences- and-Disease-895
University of Utah, Genetic Science Learning Center, Epigenetics.
http://learn.genetics.utah.edu/content/epigenetics/
Reference
Dana C. Dolinoy, Jennifer R. Weidman, Robert A. Waterland and Randy L. Jirtle. Maternal Genistein Alters Coat
Color and Protects Avy Mouse Offspring from Obesity by Modifying the Fetal Epigenome. Environ Health Perspect.114: 567–
572. 2006.
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1440782/
Paul Wade and Trevor Archer. Epigenetics: Environmental Instructions for the Genome. Environ Health
Perspect.114:A140 –A141. 2006.
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1392246/
DNA Wrap – Packaging Matters
Student Worksheet Answer Key
Step 1
Watch Chapter 1 of the video “A Tale of Two Mice,” and be prepared to discuss these questions as a class:
1. What does it mean to say that two individuals are genetically identical?
Answers may vary, but will likely include some consensus about genetically identical offspring having the same sequences of
DNA in their genes.
2. How can two genetically identical mice look so different?
Answers may vary but do not tell students the answer. The genes of genetically identical individuals are not always
expressed equally—this is known as differential gene expression.
Step 2
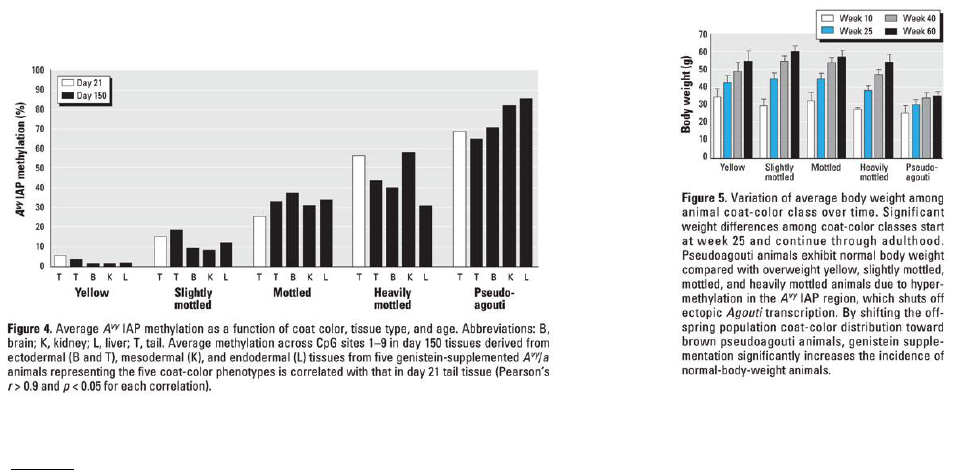
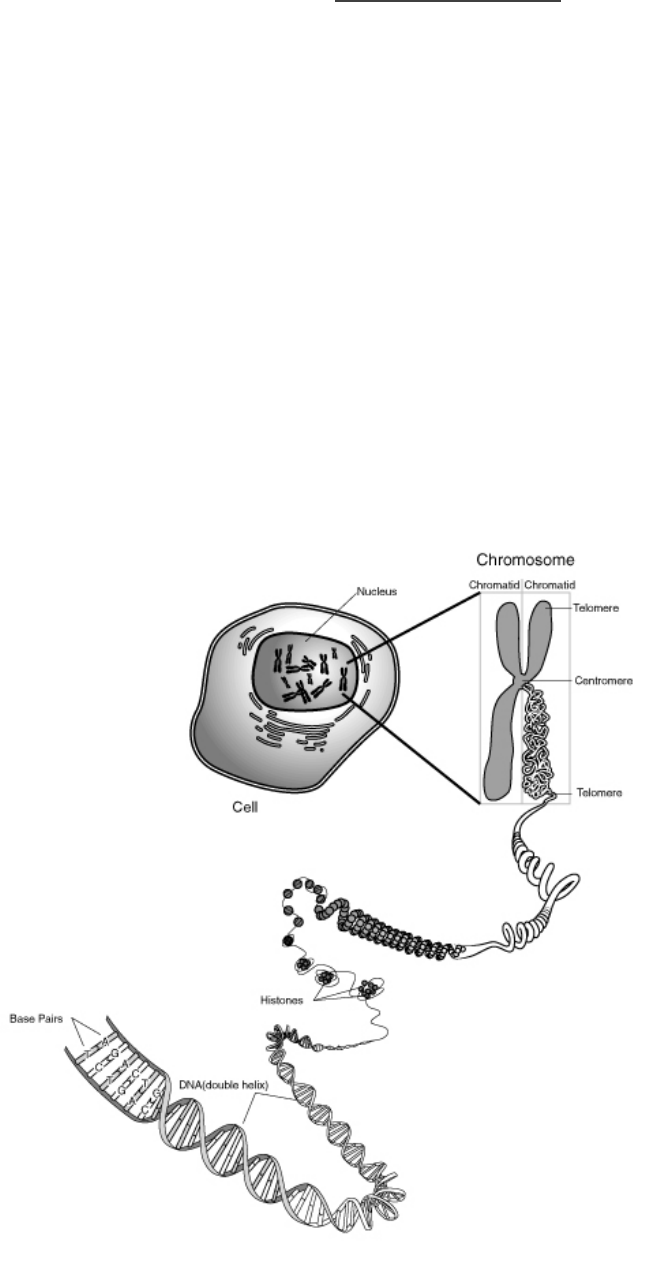
The figure below shows the location of chromosomes within a cell and the composition (i.e., DNA and
structural proteins) and three-dimensional structure of chromosomes. Individually, or with a partner, use your
textbook if necessary, to:
1. Indicate where you would find the following:
a. Condensed chromatin
b. Decondensed chromatin
c. Hydrogen bonds
d. Phosphodiester bond
e. Nucleotide
2. List the components of chromatin.
The main components are DNA and histone proteins but chromatin also includes RNA molecules and other associated proteins.
3. Describe the role of histone proteins within a chromosome.
Histone proteins act as “spools” around which DNA winds to reduce the amount of space taken up by DNA in a cell. In this
lesson, students learn that histones play both a structural and functional role in the cell since histones also play a role in gene
regulation.
Science Education Program
Step 3
Research suggests that the way DNA is “packaged” into chromatin plays an important role in genetic processes
like DNA replication, recombination, repair, and transcription. This means that changes in gene expression (i.e.,
the yellow mouse versus the brown mouse in the video you saw) can occur without changes in the DNA
structure itself (mutation).
Epigenetics is the study of other factors besides the DNA sequence that influence whether or not a gene is
transcribed into mRNA and then translated (conversion of mRNA sequence into amino acids) into a protein.
An individual’s environment, even in the womb, can influence these factors and permanently alter the
expression of genes in the adult. Alterations in epigenetic mechanisms lead to development of diseases, such as
some forms of cancer, including colorectal cancer and leukemia, neurodevelopmental disorders, obesity, and
type 2 diabetes.
Your teacher provided you with an example of an epigenetic mechanism called DNA methylation that
prevents a gene from being expressed (transcribed and translated into its protein product); this is also known as
suppression of gene expression or gene silencing.
After learning about methylation and viewing Chapter 2 of “A Tale of Two Mice,” summarize in your own
words how this epigenetic modification affects DNA structure and function.
Answers will vary, but should indicate student understanding that methylation suppresses transcription of a gene by preventing the
gene from being accessed by enzymes such as RNA polymerase that are essential for transcription to occur.
Step 4
Below are the experimental results published in Figures 4 and 5 of the Environmental Health Perspectives article
Maternal Genistein Alters Coat Color and Protects A
vy
Mouse Offspring from Obesity by Modifying the Fetal Epigenome.
In this study, pregnant mice were exposed to genistein, a component of soy, to determine if exposure to
genistein influenced gene expression among genetically identical offspring.
DNA WR APN
Look at the experimental results above (Figures 4 and 5) and complete the table on the next page
before answering the questions that follow:

Mouse Coat Color
% Methylation
in Tail Tissue (T) on day 21
(
see Figure 4
)
Body Weight (g)
at week 60
(
see Figure 5
)
Yellow
~5%
~54g
Slightly Mottled
~15%
~60g
Mottled
~36%
~58g
Heavily Mottled
~58%
~54g
Pseudo-agouti (brown)
~70%
~38g
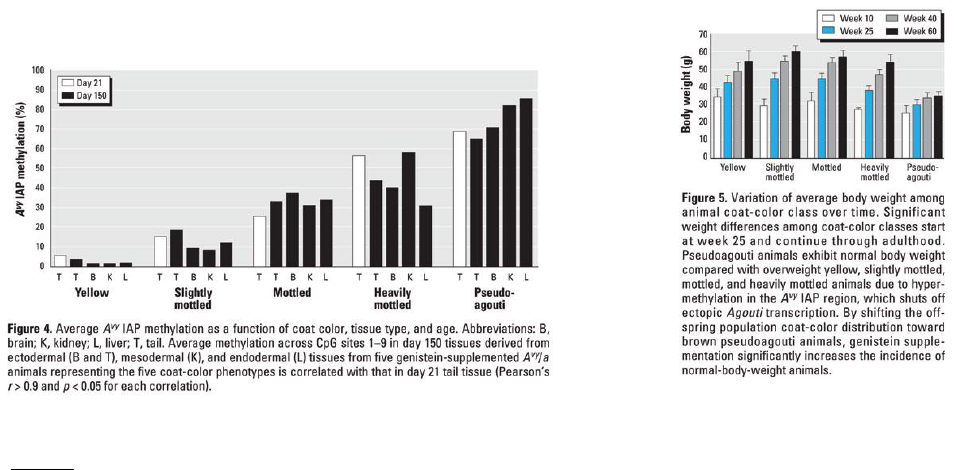
1. According to Figure 4, what is the relationship between percent methylation (y-axis) and coat color (x-axis)?
The greater the % methylation, the darker the coat color.
2. According to Figure 5, what is the relationship between body weight (y-axis) and coat color (x-axis)?
The greater the % methylation, the more protected the mouse is from obesity (mice exhibit normal body weight).
3. What explains the five different coat colors observed among offspring?
Each coat color corresponds to a different extent of methylation; the amount of methylation increases as coat color darkens.
4. In which mouse population is the Agouti gene silenced? Active? M
The Agouti gene is silenced in mice exhibiting the pseudo-agouti coat color and is most active in mice exhibiting a yellow coat.
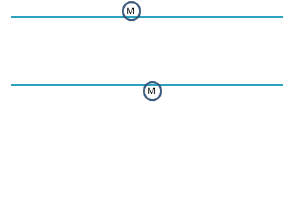
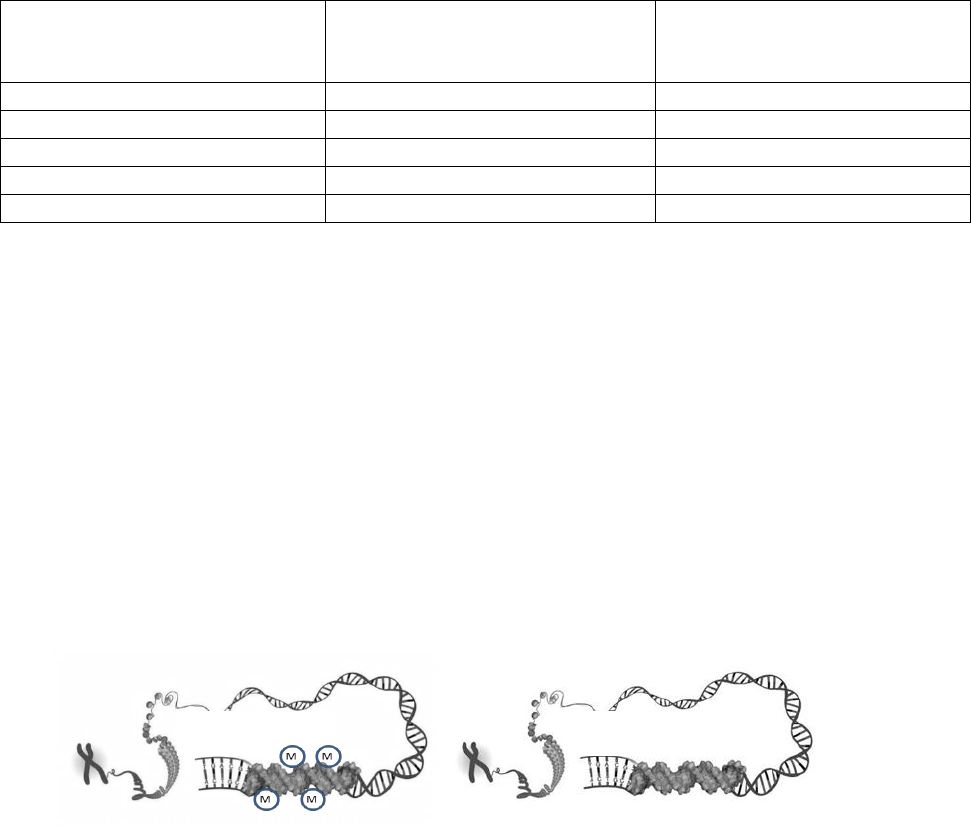
5. Assume the DNA diagrams below represent two Agouti genes, one in a brown mouse and one in a yellow
mouse. Indicate which mouse would have a methylated Agouti gene by drawing methyl groups on the
appropriate DNA strand.
The Agouti gene in the brown mouse is methylated.
Step 5
Read the authors’ conclusions below, and with a partner discuss how these conclusions could be
relevant for humans and summarize in your own words below.
“In the present study, we observed a statistically significant shift in coat-color phenotype and adult body weight distribution
among genetically identical offspring whose mothers received a diet supplemented with 250 mg/kg diet of genistein. The
shifts in coat color and body weight were mediated by increased methylation … of the Agouti gene. Hypermethylation in
the genistein-supplemented population results in decreased ectopic Agouti expression, which reduces yellow phaeomelanin
production and protects against adult-onset obesity. This is the first study to demonstrate that exposure to dietary genistein
in utero, at levels present in human adult populations consuming high-soy diets, affects coat color and reduces population
incidence of obesity by altering the epigenome in mice. Thus, an active ingredient in soy enhances the early establishment
of DNA methylation.… [O]ur findings show that early nutritional and environmentally induced epigenetic modifications
can provide an alternative mechanism for varying individual susceptibilities to environmental agents. Our results also suggest
a plausible explanation for the lower incidence of certain cancers in Asians compared with Westerners (Chang et al. 2001;
Lee et al. 1991) as well as the increased cancer incidence in Asians who immigrate to the United States (Ziegler et al.
1993).”TT
Answers will vary, but students should conclude that these results may also apply to humans, and that diet, especially during pregnancy,
can influence our epigenome and ultimately determine our susceptibility to cancer and disease. This example also clearly demonstrates
that DNA methylation does not always lead to negative consequences for the individual.
Brown Mouse
Yellow Mouse
DNA Wrap – Packaging Matters
Student Worksheet
Name:
Step 1
Watch Chapter 1 of the video “A Tale of Two Mice,” and be prepared to discuss these questions as a class:
3. What does it mean to say that two individuals are genetically identical?
4. How can two genetically identical mice look so different?
Step 2
The figure below shows the location of chromosomes within a cell and the composition (i.e., DNA and
structural proteins) and three-dimensional structure of chromosomes. Individually, or with a partner, use your
textbook if necessary, to:
1. Indicate where you would find the following:
a. Condensed chromatin
b. Decondensed chromatin
c. Hydrogen bonds
d. Phosphodiester bond
e. Nucleotide
3. List the components of chromatin.
3. Describe the role of histone proteins within a chromosome.
WRAP– Science Education Program
Source: http://commons.wikimedia.org/wiki/File:Chromosome.gif
Step 3
Research suggests that the way DNA is “packaged” into chromatin plays an important role in genetic
processes like DNA replication, recombination, repair, and transcription. This means that changes in gene
expression (i.e., the yellow mouse versus the brown mouse in the video you saw) can occur without changes in the DNA structure
itself (mutation).
Epigenetics is the study of other factors besides the DNA sequence that influence whether or not a gene is
transcribed into mRNA and then translated (conversion of mRNA sequence into amino acids) into a protein.
An individual’s environment, even in the womb, can influence these factors and permanently alter the
expression of genes in the adult. Alterations in epigenetic mechanisms lead to development of diseases, such
as some forms of cancer, including colorectal cancer and leukemia, neurodevelopmental disorders, obesity,
and type 2 diabetes.
Your teacher provided you with an example of an epigenetic mechanism called DNA methylation that
prevents a gene from being expressed (transcribed and translated into its protein product); this is also known
as suppression of gene expression or gene silencing.
After learning about methylation and viewing Chapter 2 of “A Tale of Two Mice,” summarize in your own
words how this epigenetic modification affects DNA structure and function.
Step 4
Below are the experimental results published in Figures 4 and 5 of the Environmental Health Perspectives article
Maternal Genistein Alters Coat Color and Protects A
vy
Mouse Offspring from Obesity by Modifying the Fetal Epigenome.
In this study, pregnant mice were exposed to genistein, a component of soy, to determine if exposure to
genistein influenced gene expression among genetically identical offspring.
DNA WR APN
Look at the experimental results above (Figures 4 and 5) and complete the table on the next page
before answering the questions that follow:

Mouse Coat Color
% Methylation
in Tail Tissue (T) on day 21
(
see Figure 4
)
Body Weight (g)
at week 60
(
see Figure 5
)
Yellow
Slightly Mottled
Mottled
Heavily Mottled
Pseudo-agouti (brown)
1. According to Figure 4, what is the relationship between percent methylation (y-axis) and coat color (x-
axis)?
2. According to Figure 5, what is the relationship between body weight (y-axis) and coat color (x-axis)?
3. What explains the five different coat colors observed among offspring?
4. In which mouse population is the Agouti gene silenced? Active? M
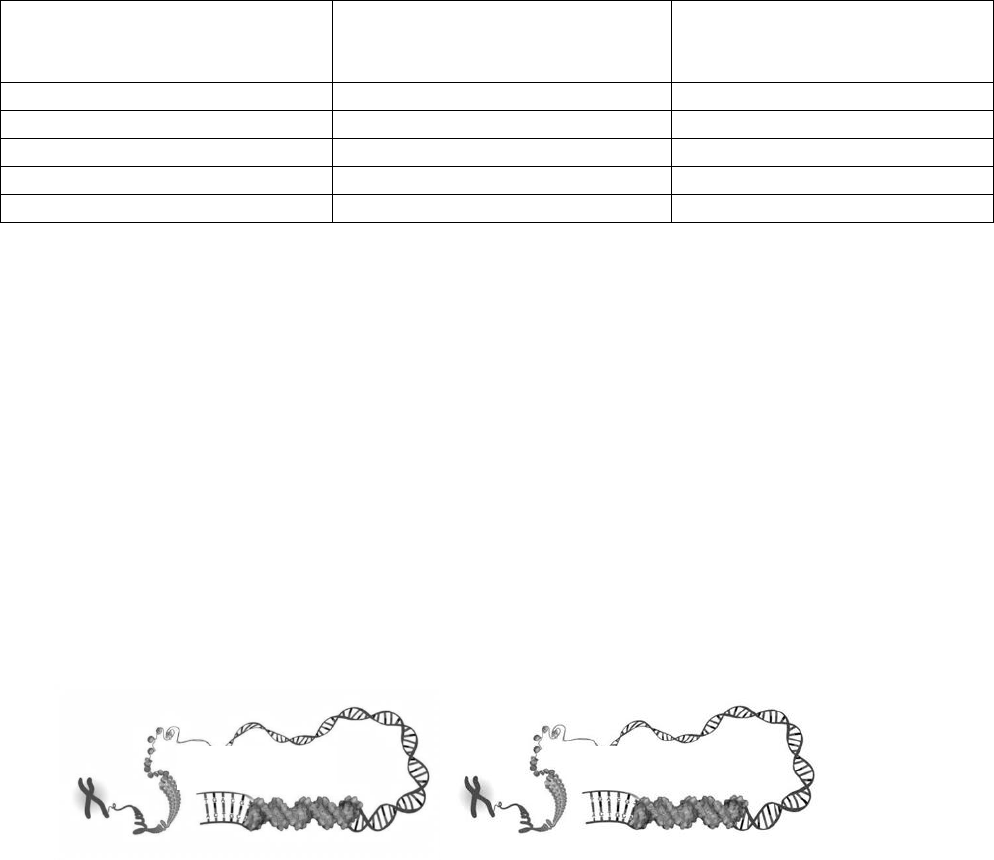
6. Assume the DNA diagrams below represent two Agouti genes, one in a brown mouse and one in a yellow
mouse. Indicate which mouse would have a methylated Agouti gene by drawing methyl groups on
the appropriate DNA molecule below.
Step 5
Read the authors’ conclusions below, and with a partner discuss how these conclusions could be
relevant for humans and summarize in your own words below.
“In the present study, we observed a statistically significant shift in coat-color phenotype and adult body weight
distribution among genetically identical offspring whose mothers received a diet supplemented with 250 mg/kg
diet of genistein. The shifts in coat color and body weight were mediated by increased methylation … of the
Agouti gene. Hypermethylation in the genistein-supplemented population results in decreased ectopic Agouti
expression, which reduces yellow phaeomelanin production and protects against adult-onset obesity. This is the
first study to demonstrate that exposure to dietary genistein in utero, at levels present in human adult populations
consuming high-soy diets, affects coat color and reduces population incidence of obesity by altering the
epigenome in mice. Thus, an active ingredient in soy enhances the early establishment of DNA methylation.…
[O]ur findings show that early nutritional and environmentally induced epigenetic modifications can provide
an alternative mechanism for varying individual susceptibilities to environmental agents. Our results also suggest
a plausible explanation for the lower incidence of certain cancers in Asians compared with Westerners (Chang
et al. 2001; Lee et al. 1991) as well as the increased cancer incidence in Asians who immigrate to the United
States (Ziegler et al. 1993).”TT
Brown Mouse
Yellow Mouse
